MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
- PMID: 17694064
- PMCID: PMC3857099
- DOI: 10.1038/nmeth1079
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
Abstract
MicroRNAs are predicted to regulate thousands of mammalian genes, but relatively few targets have been experimentally validated and few microRNA loss-of-function phenotypes have been assigned. As an alternative to chemically modified antisense oligonucleotides, we developed microRNA inhibitors that can be expressed in cells, as RNAs produced from transgenes. Termed 'microRNA sponges', these competitive inhibitors are transcripts expressed from strong promoters, containing multiple, tandem binding sites to a microRNA of interest. When vectors encoding these sponges are transiently transfected into cultured cells, sponges derepress microRNA targets at least as strongly as chemically modified antisense oligonucleotides. They specifically inhibit microRNAs with a complementary heptameric seed, such that a single sponge can be used to block an entire microRNA seed family. RNA polymerase II promoter (Pol II)-driven sponges contain a fluorescence reporter gene for identification and sorting of sponge-treated cells. We envision the use of stably expressed sponges in animal models of disease and development.
Conflict of interest statement
The authors declare no competing financial interests.
Figures
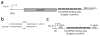
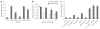
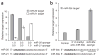
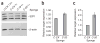
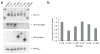
Comment in
-
Soaking up small RNAs.Nat Methods. 2007 Sep;4(9):694-5. doi: 10.1038/nmeth0907-694. Nat Methods. 2007. PMID: 17762876 No abstract available.
References
-
- Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20. - PubMed
-
- Lu J, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–838. - PubMed
-
- Li QJ, et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell. 2007;129:147–161. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

