The roles of binding site arrangement and combinatorial targeting in microRNA repression of gene expression
- PMID: 17697356
- PMCID: PMC2374997
- DOI: 10.1186/gb-2007-8-8-r166
The roles of binding site arrangement and combinatorial targeting in microRNA repression of gene expression
Abstract
Background: MicroRNAs (miRNAs) are small noncoding RNAs that bind mRNA target transcripts and repress gene expression. They have been implicated in multiple diseases, such as cancer, but the mechanisms of this involvement are not well understood. Given the complexity and degree of interactions between miRNAs and target genes, understanding how miRNAs achieve their specificity is important to understanding miRNA function and identifying their role in disease.
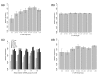
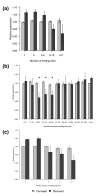
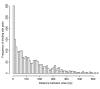
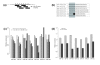
Results: Here we report factors that influence miRNA regulation by considering the effects of both single and multiple miRNAs targeting human genes. In the case of single miRNA targeting, we developed a metric that integrates miRNA and mRNA expression data to calculate how changes in miRNA expression affect target mRNA expression. Using the metric, our global analysis shows that the repression of a given miRNA on a target mRNA is modulated by 3' untranslated region length, the number of target sites, and the distance between a pair of binding sites. Additionally, we show that some miRNAs preferentially repress transcripts with longer CTG repeats, suggesting a possible role for miRNAs in repeat expansion disorders such as myotonic dystrophy. We also examine the large class of genes targeted by multiple miRNAs and show that specific types of genes are progressively more enriched as the number of targeting miRNAs increases. Expression microarray data further show that these highly targeted genes are downregulated relative to genes targeted by few miRNAs, which suggests that highly targeted genes are tightly regulated and that their dysregulation may lead to disease. In support of this idea, cancer genes are strongly enriched among highly targeted genes.
Conclusion: Our data show that the rules governing miRNA targeting are complex, but that understanding the mechanisms that drive such control can uncover miRNAs' role in disease. Our study suggests that the number and arrangement of miRNA recognition sites can influence the degree and specificity of miRNA-mediated gene repression.
Figures

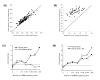
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

