A role for yeast and human translesion synthesis DNA polymerases in promoting replication through 3-methyl adenine
- PMID: 17698580
- PMCID: PMC2168906
- DOI: 10.1128/MCB.01079-07
A role for yeast and human translesion synthesis DNA polymerases in promoting replication through 3-methyl adenine
Abstract
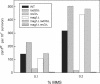
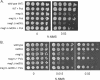
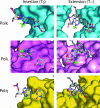
3-Methyl adenine (3meA), a minor-groove DNA lesion, presents a strong block to synthesis by replicative DNA polymerases (Pols). To elucidate the means by which replication through this DNA lesion is mediated in eukaryotic cells, here we carry out genetic studies in the yeast Saccharomyces cerevisiae treated with the alkylating agent methyl methanesulfonate. From the studies presented here, we infer that replication through the 3meA lesion in yeast cells can be mediated by the action of three Rad6-Rad18-dependent pathways that include translesion synthesis (TLS) by Pol(eta) or -zeta and an Mms2-Ubc13-Rad5-dependent pathway which presumably operates via template switching. We also express human Pols iota and kappa in yeast cells and show that they too can mediate replication through the 3meA lesion in yeast cells, indicating a high degree of evolutionary conservation of the mechanisms that control TLS in yeast and human cells. We discuss these results in the context of previous observations that have been made for the roles of Pols eta, iota, and kappa in promoting replication through the minor-groove N2-dG adducts.
Figures
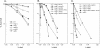
Similar articles
-
Mms2-Ubc13-dependent and -independent roles of Rad5 ubiquitin ligase in postreplication repair and translesion DNA synthesis in Saccharomyces cerevisiae.Mol Cell Biol. 2006 Oct;26(20):7783-90. doi: 10.1128/MCB.01260-06. Epub 2006 Aug 14. Mol Cell Biol. 2006. PMID: 16908531 Free PMC article.
-
Requirement of RAD5 and MMS2 for postreplication repair of UV-damaged DNA in Saccharomyces cerevisiae.Mol Cell Biol. 2002 Apr;22(7):2419-26. doi: 10.1128/MCB.22.7.2419-2426.2002. Mol Cell Biol. 2002. PMID: 11884624 Free PMC article.
-
Requirement of Rad5 for DNA polymerase zeta-dependent translesion synthesis in Saccharomyces cerevisiae.Genetics. 2008 Sep;180(1):73-82. doi: 10.1534/genetics.108.091066. Epub 2008 Aug 30. Genetics. 2008. PMID: 18757916 Free PMC article.
-
Regulation of DNA damage tolerance in mammalian cells by post-translational modifications of PCNA.Mutat Res. 2017 Oct;803-805:82-88. doi: 10.1016/j.mrfmmm.2017.06.004. Epub 2017 Jun 21. Mutat Res. 2017. PMID: 28666590 Review.
-
Role of yeast Rad5 and its human orthologs, HLTF and SHPRH in DNA damage tolerance.DNA Repair (Amst). 2010 Mar 2;9(3):257-67. doi: 10.1016/j.dnarep.2009.12.013. Epub 2010 Jan 21. DNA Repair (Amst). 2010. PMID: 20096653 Review.
Cited by
-
Polymerase theta-mediated end joining of replication-associated DNA breaks in C. elegans.Genome Res. 2014 Jun;24(6):954-62. doi: 10.1101/gr.170431.113. Epub 2014 Mar 10. Genome Res. 2014. PMID: 24614976 Free PMC article.
-
DNA repair mechanisms and the bypass of DNA damage in Saccharomyces cerevisiae.Genetics. 2013 Apr;193(4):1025-64. doi: 10.1534/genetics.112.145219. Genetics. 2013. PMID: 23547164 Free PMC article. Review.
-
Involvement of Rev1 in alkylating agent-induced loss of heterozygosity in Oryzias latipes.Genes Cells. 2020 Feb;25(2):124-138. doi: 10.1111/gtc.12746. Epub 2020 Feb 5. Genes Cells. 2020. PMID: 31917895 Free PMC article.
-
Acetylation of Werner syndrome protein (WRN): relationships with DNA damage, DNA replication and DNA metabolic activities.Biogerontology. 2014 Aug;15(4):347-66. doi: 10.1007/s10522-014-9506-3. Epub 2014 Jun 26. Biogerontology. 2014. PMID: 24965941 Free PMC article.
-
Contributions of DNA repair and damage response pathways to the non-linear genotoxic responses of alkylating agents.Mutat Res Rev Mutat Res. 2016 Jan-Mar;767:77-91. doi: 10.1016/j.mrrev.2015.11.001. Epub 2015 Dec 2. Mutat Res Rev Mutat Res. 2016. PMID: 27036068 Free PMC article. Review.
References
-
- Beranek, D. T., C. C. Weis, and D. H. Swenson. 1980. A comprehensive quantitative analysis of methylated and ethylated DNA using high pressure liquid chromatography. Carcinogenesis 1:595-606. - PubMed
-
- Bjoras, M., A. Klungland, R. F. Johansen, and E. Seeberg. 1995. Purification and properties of the alkylation repair DNA glycosylase encoded MAG gene from Saccharomyces cerevisiae. Biochemistry 34:4577-4582. - PubMed
-
- de los Santos, C., T. Zaliznyak, and F. Johnson. 2001. NMR characterization of a DNA duplex containing the major acrolein-derived deoxyguanosine adduct γ-OH-1,-N2-propano-2′-deoxyguanosine. J. Biol. Chem. 276:9077-9082. - PubMed
-
- Engelward, B. P., J. M. Allan, A. J. Dreslin, J. D. Kelly, M. M. Wu, B. Gold, and L. D. Samson. 1998. A chemical and genetic approach together define the biological consequences of 3-methylanine lesions in the mammalian genome. J. Biol. Chem. 273:5412-5418. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
