Genetic characteristics and clonal dissemination of beta-lactamase-negative ampicillin-resistant Haemophilus influenzae strains isolated from the upper respiratory tract of patients in Japan
- PMID: 17698631
- PMCID: PMC2151452
- DOI: 10.1128/AAC.00422-07
Genetic characteristics and clonal dissemination of beta-lactamase-negative ampicillin-resistant Haemophilus influenzae strains isolated from the upper respiratory tract of patients in Japan
Abstract
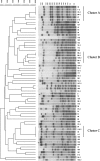
We evaluated the recent prevalence of antimicrobial-resistant Haemophilus influenzae isolated from the upper respiratory tracts (URT) of patients in Japan. Mutations in the ftsI gene, which encodes penicillin binding protein 3 (PBP3), and the clonal dissemination of the resistant strains were also investigated. A total of 264 H. influenzae isolates were collected from patients with URT infections. According to the criteria of the Clinical and Laboratory Standards Institute for the susceptibility of H. influenzae to ampicillin (AMP), the isolates were distributed as follows: 161 (61.0%) susceptible strains (MIC < or = 1 microg/ml), 37 (14.0%) intermediately resistant strains (MIC = 2 microg/ml), and 66 (25.0%) resistant strains (MIC > or = 4 microg/ml). According to PCR-based genotyping, 172 (65.1%) of the isolates had mutations in the ftsI gene and were negative for the beta-lactamase (bla) gene. These 172 isolates were thus defined as genetically beta-lactamase-negative ampicillin-resistant (gBLNAR) strains. The ftsI mutant group included 98 (37.1%) strains with group I/II mutations in the variable mutated region (group I/II gBLNAR) and 74 (28.0%) strains with group III mutations in the highly mutated region (group III gBLNAR). Eighty-seven (33.0%) of the isolates were genetically beta-lactamase-negative ampicillin-susceptible (gBLNAS) strains. The group III gBLNAR strains showed resistance to beta-lactams. Only five strains (1.9%) were positive for a bla gene encoding TEM-type beta-lactamase. The three clusters consisting of 16 strains found among the 61 BLNAR strains (MIC > or = 4 microg/ml and without the bla gene) showed identical or closely related DNA restriction fragment patterns. Those isolates were frequently identified among strains with a MIC to AMP of 16 microg/ml. The current study demonstrates the apparent dissemination and spread of a resistant clone of H. influenzae among medical centers in Japan. The gBLNAR strains show a remarkable prevalence among H. influenzae isolates, with the prevalence increasing with time. This fact should be taken into account when treating URT infections.
Figures

Similar articles
-
Antimicrobial resistance in Haemophilus influenzae isolated from the nasopharynx among Japanese children with acute otitis media.Acta Otolaryngol. 2006 Feb;126(2):130-7. doi: 10.1080/00016480500312455. Acta Otolaryngol. 2006. PMID: 16428188
-
Mutant ftsI genes in the emergence of penicillin-binding protein-mediated beta-lactam resistance in Haemophilus influenzae in Norway.Clin Microbiol Infect. 2010 Aug;16(8):1117-24. doi: 10.1111/j.1469-0691.2009.03052.x. Epub 2009 Sep 8. Clin Microbiol Infect. 2010. PMID: 19737286
-
Antimicrobial resistance of Haemophilus influenzae isolated from the nasopharynx of Japanese children with acute otitis media.Acta Otolaryngol. 2006 Mar;126(3):240-7. doi: 10.1080/00016480500314287. Acta Otolaryngol. 2006. PMID: 16618648
-
[Beta-lactamase negative ampicillin-resistant Haemophilus influenzae(BLNAR)].Nihon Rinsho. 2001 Apr;59(4):688-93. Nihon Rinsho. 2001. PMID: 11304990 Review. Japanese.
-
Revisiting mutational resistance to ampicillin and cefotaxime in Haemophilus influenzae.Genome Med. 2024 Dec 4;16(1):140. doi: 10.1186/s13073-024-01406-4. Genome Med. 2024. PMID: 39633433 Free PMC article.
Cited by
-
Characteristics of Haemophilus influenzae invasive isolates from Portugal following routine childhood vaccination against H. influenzae serotype b (2002-2010).Eur J Clin Microbiol Infect Dis. 2014 Apr;33(4):603-10. doi: 10.1007/s10096-013-1994-6. Epub 2013 Oct 24. Eur J Clin Microbiol Infect Dis. 2014. PMID: 24154654
-
Nontypable Haemophilus influenzae Septicemia and Urinary Tract Infection Associated with Renal Stone Disease.Open Microbiol J. 2018 Jul 31;12:243-247. doi: 10.2174/1874285801812010243. eCollection 2018. Open Microbiol J. 2018. PMID: 30197697 Free PMC article.
-
Multilocus sequence typing and ftsI sequencing: a powerful tool for surveillance of penicillin-binding protein 3-mediated beta-lactam resistance in nontypeable Haemophilus influenzae.BMC Microbiol. 2014 May 20;14:131. doi: 10.1186/1471-2180-14-131. BMC Microbiol. 2014. PMID: 24884375 Free PMC article.
-
Incidence survey of acute otitis media in children in Sado Island, Japan--Sado Otitis Media Study (SADOMS).PLoS One. 2013 Jul 2;8(7):e68711. doi: 10.1371/journal.pone.0068711. Print 2013. PLoS One. 2013. PMID: 23844235 Free PMC article.
-
Multiclonal Expansion and High Prevalence of β-Lactamase-Negative Haemophilus influenzae with High-Level Ampicillin Resistance in Japan and Susceptibility to Quinolones.Antimicrob Agents Chemother. 2018 Aug 27;62(9):e00851-18. doi: 10.1128/AAC.00851-18. Print 2018 Sep. Antimicrob Agents Chemother. 2018. PMID: 29987153 Free PMC article.
References
-
- Clairoux, N., M. Picard, A. Brochu, N. Rousseau, P. Gourde, D. Beauchamp, T. R. Parr, Jr., M. G. Bergeron, and F. Malouin. 1992. Molecular Basis of non-β-lactamase-mediated resistance to β-lactam antibiotics in strains of Haemophilus influenzae. Antimicrob. Agents Chemother. 36:1504-1513. - PMC - PubMed
-
- Clinical and Laboratory Standards Institute. 2006. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically, M7-A7. Approved standard, 7th ed. Clinical and Laboratory Standards Institute, Wayne, PA.
-
- Dabernat, H., P. Geslin, F. Megraud, P. Begue, J. Bouleteix, C. Dubreuil, F. de La Roque, A. Trinh, and A. Scheimberg. 1998. Effect of cefixime or co-amoxiclav treatment on nasopharyngeal carriage of Streptococcus pneumoniae and Haemophilus influenzae in children with acute otitis media. J. Antimoicrob. Chemother. 41:253-258. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

