von Hippel Lindau binding protein 1-mediated degradation of integrase affects HIV-1 gene expression at a postintegration step
- PMID: 17698809
- PMCID: PMC1959430
- DOI: 10.1073/pnas.0705162104
von Hippel Lindau binding protein 1-mediated degradation of integrase affects HIV-1 gene expression at a postintegration step
Abstract
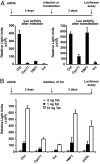
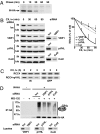
HIV-1 integrase, the viral enzyme responsible for provirus integration into the host genome, can be actively degraded by the ubiquitin-proteasome pathway. Here, we identify von Hippel-Lindau binding protein 1(VBP1), a subunit of the prefoldin chaperone, as an integrase cellular binding protein that bridges interaction between integrase and the cullin2 (Cul2)-based von Hippel-Lindau (VHL) ubiquitin ligase. We demonstrate that VBP1 and Cul2/VHL are required for proper HIV-1 expression at a step between integrase-dependent proviral integration into the host genome and transcription of viral genes. Using both an siRNA approach and Cul2/VHL mutant cells, we show that VBP1 and the Cul2/VHL ligase cooperate in the efficient polyubiquitylation of integrase and its subsequent proteasome-mediated degradation. Results presented here support a role for integrase degradation by the prefoldin-VHL-proteasome pathway in the integration-transcription transition of the viral replication cycle.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

