ErpA, an iron sulfur (Fe S) protein of the A-type essential for respiratory metabolism in Escherichia coli
- PMID: 17698959
- PMCID: PMC1959432
- DOI: 10.1073/pnas.0705829104
ErpA, an iron sulfur (Fe S) protein of the A-type essential for respiratory metabolism in Escherichia coli
Abstract
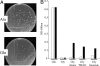
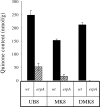
Understanding the biogenesis of iron-sulfur (Fe-S) proteins is relevant to many fields, including bioenergetics, gene regulation, and cancer research. Several multiprotein complexes assisting Fe-S assembly have been identified in both prokaryotes and eukaryotes. Here, we identify in Escherichia coli an A-type Fe-S protein that we named ErpA. Remarkably, erpA was found essential for growth of E. coli in the presence of oxygen or alternative electron acceptors. It was concluded that isoprenoid biosynthesis was impaired by the erpA mutation. First, the eukaryotic mevalonate-dependent pathway for biosynthesis of isopentenyl diphosphate restored the respiratory defects of an erpA mutant. Second, the erpA mutant contained a greatly reduced amount of ubiquinone and menaquinone. Third, ErpA bound Fe-S clusters and transferred them to apo-IspG, a protein catalyzing isopentenyl diphosphate biosynthesis in E. coli. Surprisingly, the erpA gene maps at a distance from any other Fe-S biogenesis-related gene. ErpA is an A-type Fe-S protein that is characterized by an essential role in cellular metabolism.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
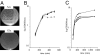
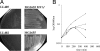

Similar articles
-
A-Type Carrier Proteins Are Involved in [4Fe-4S] Cluster Insertion into the Radical S-Adenosylmethionine Protein MoaA for the Synthesis of Active Molybdoenzymes.J Bacteriol. 2021 May 20;203(12):e0008621. doi: 10.1128/JB.00086-21. Epub 2021 Mar 29. J Bacteriol. 2021. PMID: 33782054 Free PMC article.
-
A Regulatory Circuit Composed of a Transcription Factor, IscR, and a Regulatory RNA, RyhB, Controls Fe-S Cluster Delivery.mBio. 2016 Sep 20;7(5):e00966-16. doi: 10.1128/mBio.00966-16. mBio. 2016. PMID: 27651365 Free PMC article.
-
Escherichia coli monothiol glutaredoxin GrxD replenishes Fe-S clusters to the essential ErpA A-type carrier under low iron stress.J Biol Chem. 2024 Aug;300(8):107506. doi: 10.1016/j.jbc.2024.107506. Epub 2024 Jun 27. J Biol Chem. 2024. PMID: 38944118 Free PMC article.
-
Assembly of Fe/S proteins in bacterial systems: Biochemistry of the bacterial ISC system.Biochim Biophys Acta. 2015 Jun;1853(6):1436-47. doi: 10.1016/j.bbamcr.2014.12.009. Epub 2014 Dec 13. Biochim Biophys Acta. 2015. PMID: 25510311 Review.
-
Structure, function, and formation of biological iron-sulfur clusters.Annu Rev Biochem. 2005;74:247-81. doi: 10.1146/annurev.biochem.74.082803.133518. Annu Rev Biochem. 2005. PMID: 15952888 Review.
Cited by
-
A-Type Carrier Proteins Are Involved in [4Fe-4S] Cluster Insertion into the Radical S-Adenosylmethionine Protein MoaA for the Synthesis of Active Molybdoenzymes.J Bacteriol. 2021 May 20;203(12):e0008621. doi: 10.1128/JB.00086-21. Epub 2021 Mar 29. J Bacteriol. 2021. PMID: 33782054 Free PMC article.
-
IscR of Rhodobacter sphaeroides functions as repressor of genes for iron-sulfur metabolism and represents a new type of iron-sulfur-binding protein.Microbiologyopen. 2015 Oct;4(5):790-802. doi: 10.1002/mbo3.279. Epub 2015 Aug 1. Microbiologyopen. 2015. PMID: 26235649 Free PMC article.
-
Metal utilization in genome-reduced bacteria: Do human mycoplasmas rely on iron?Comput Struct Biotechnol J. 2021 Oct 18;19:5752-5761. doi: 10.1016/j.csbj.2021.10.022. eCollection 2021. Comput Struct Biotechnol J. 2021. PMID: 34765092 Free PMC article. Review.
-
Functional replacement of isoprenoid pathways in Rhodobacter sphaeroides.Microb Biotechnol. 2020 Jul;13(4):1082-1093. doi: 10.1111/1751-7915.13562. Epub 2020 Mar 24. Microb Biotechnol. 2020. PMID: 32207882 Free PMC article.
-
Salmonella enterica requires ApbC function for growth on tricarballylate: evidence of functional redundancy between ApbC and IscU.J Bacteriol. 2008 Jul;190(13):4596-602. doi: 10.1128/JB.00262-08. Epub 2008 Apr 25. J Bacteriol. 2008. PMID: 18441067 Free PMC article.
References
-
- Lill R, Mühlenhoff U. Annu Rev Cell Dev Biol. 2006;22:457–486. - PubMed
-
- Barras F, Loiseau L, Py B. Adv Microb Physiol. 2005;50:41–101. - PubMed
-
- Fontecave M, Ollagnier-de Choudens S, Py B, Barras F. J Biol Inorg Chem. 2005;10:713–721. - PubMed
-
- Johnson DC, Dean DR, Smith AD, Johnson MK. Annu Rev Biochem. 2005;74:247–281. - PubMed
-
- Beinert H, Holm RH, Munck E. Science. 1997;277:653–659. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

