Subtypes of the plasmid-encoded serine protease EspP in Shiga toxin-producing Escherichia coli: distribution, secretion, and proteolytic activity
- PMID: 17704265
- PMCID: PMC2075056
- DOI: 10.1128/AEM.00920-07
Subtypes of the plasmid-encoded serine protease EspP in Shiga toxin-producing Escherichia coli: distribution, secretion, and proteolytic activity
Abstract
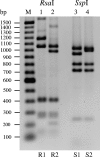
We investigated the prevalence, distribution, and structure of espP in Shiga toxin-producing Escherichia coli (STEC) and assessed the secretion and proteolytic activity of the encoded autotransporter protein EspP (extracellular serine protease, plasmid encoded). espP was identified in 56 of 107 different STEC serotypes. Sequencing of a 3,747-bp region of the 3,900-bp espP gene distinguished four alleles (espPalpha, espPbeta, espPgamma, and espPdelta), with 99.9%, 99.2%, 95.3%, and 95.1% homology, respectively, to espP of E. coli O157:H7 strain EDL933. The espPbeta, espPgamma, and espPdelta genes contained unique insertions and/or clustered point mutations that enabled allele-specific PCRs; these demonstrated the presence of espPalpha, espPbeta, espPgamma, and espPdelta in STEC isolates belonging to 17, 16, 15, and 8 serotypes, respectively. Among four subtypes of EspP encoded by these alleles, EspPalpha (produced by enterohemorrhagic E. coli [EHEC] O157:H7 and the major non-O157 EHEC serotypes) and EspPgamma cleaved pepsin A, human coagulation factor V, and an oligopeptide alanine-alanine-proline-leucine-para-nitroaniline, whereas EspPbeta and EspPdelta either were not secreted or were proteolytically inactive. The lack of proteolysis correlated with point mutations near the active serine protease site. We conclude that espP is widely distributed among STEC strains and displays genetic heterogeneity, which can be used for subtyping and which affects EspP activity. The presence of proteolytically active EspP in EHEC serogroups O157, O26, O111, and O145, which are bona fide human pathogens, suggests that EspP might play a role as an EHEC virulence factor.
Figures


Similar articles
-
Structure and function relationship of the autotransport and proteolytic activity of EspP from Shiga toxin-producing Escherichia coli.PLoS One. 2009 Jul 1;4(7):e6100. doi: 10.1371/journal.pone.0006100. PLoS One. 2009. PMID: 19568421 Free PMC article.
-
Prevalence, biogenesis, and functionality of the serine protease autotransporter EspP.Toxins (Basel). 2012 Dec 28;5(1):25-48. doi: 10.3390/toxins5010025. Toxins (Basel). 2012. PMID: 23274272 Free PMC article. Review.
-
Serine protease espP subtype alpha, but not beta or gamma, of Shiga toxin-producing Escherichia coli is associated with highly pathogenic serogroups.Int J Med Microbiol. 2009 Apr;299(4):247-54. doi: 10.1016/j.ijmm.2008.08.006. Epub 2008 Nov 25. Int J Med Microbiol. 2009. PMID: 19036636
-
DNA sequence and analysis of a 90.1-kb plasmid in Shiga toxin-producing Escherichia coli (STEC) O145:NM 83-75.Plasmid. 2012 Jul;68(1):25-32. doi: 10.1016/j.plasmid.2012.02.001. Epub 2012 Feb 19. Plasmid. 2012. PMID: 22370037
-
Enteroaggregative Escherichia coli plasmid-encoded toxin.Future Microbiol. 2010 Jul;5(7):1005-13. doi: 10.2217/fmb.10.69. Future Microbiol. 2010. PMID: 20632801 Review.
Cited by
-
Enterohemorrhagic E. coli (EHEC)-Secreted Serine Protease EspP Stimulates Electrogenic Ion Transport in Human Colonoid Monolayers.Toxins (Basel). 2018 Sep 1;10(9):351. doi: 10.3390/toxins10090351. Toxins (Basel). 2018. PMID: 30200426 Free PMC article.
-
Serine Protease Autotransporters of the Enterobacteriaceae (SPATEs): Out and About and Chopping It Up.Microorganisms. 2019 Nov 21;7(12):594. doi: 10.3390/microorganisms7120594. Microorganisms. 2019. PMID: 31766493 Free PMC article. Review.
-
Structure and function relationship of the autotransport and proteolytic activity of EspP from Shiga toxin-producing Escherichia coli.PLoS One. 2009 Jul 1;4(7):e6100. doi: 10.1371/journal.pone.0006100. PLoS One. 2009. PMID: 19568421 Free PMC article.
-
Crystal structure of the passenger domain of the Escherichia coli autotransporter EspP.J Mol Biol. 2011 Nov 11;413(5):985-1000. doi: 10.1016/j.jmb.2011.09.028. Epub 2011 Sep 22. J Mol Biol. 2011. PMID: 21964244 Free PMC article.
-
Prevalence, biogenesis, and functionality of the serine protease autotransporter EspP.Toxins (Basel). 2012 Dec 28;5(1):25-48. doi: 10.3390/toxins5010025. Toxins (Basel). 2012. PMID: 23274272 Free PMC article. Review.
References
-
- Bandelt, H.-J., and A. W. M. Dress. 1992. Split decomposition: a new and useful approach to phylogenetic analysis of distance data. Mol. Phylogenet. Evol. 1:242-252. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

