Genomic mapping of Suppressor of Hairy-wing binding sites in Drosophila
- PMID: 17705839
- PMCID: PMC2374998
- DOI: 10.1186/gb-2007-8-8-r167
Genomic mapping of Suppressor of Hairy-wing binding sites in Drosophila
Abstract
Background: Insulator elements are proposed to play a key role in the organization of the regulatory architecture of the genome. In Drosophila, one of the best studied is the gypsy retrotransposon insulator, which is bound by the Suppressor of Hairy-wing (Su [Hw]) transcriptional regulator. Immunolocalization studies suggest that there are several hundred Su(Hw) sites in the genome, but few of these endogenous Su(Hw) binding sites have been identified.
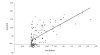
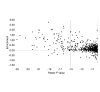
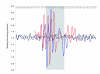
Results: We used chromatin immunopurification with genomic microarray analysis to identify in vivo Su(Hw) binding sites across the 3 megabase Adh region. We find 60 sites, and these enabled the construction of a robust new Su(Hw) binding site consensus. In contrast to the gypsy insulator, which contains tightly clustered Su(Hw) binding sites, endogenous sites generally occur as isolated sites. These endogenous sites have three key features. In contrast to most analyses of DNA-binding protein specificity, we find that strong matches to the binding consensus are good predictors of binding site occupancy. Examination of occupancy in different tissues and developmental stages reveals that most Su(Hw) sites, if not all, are constitutively occupied, and these isolated Su(Hw) sites are generally highly conserved. Analysis of transcript levels in su(Hw) mutants indicate widespread and general changes in gene expression. Importantly, the vast majority of genes with altered expression are not associated with clustering of Su(Hw) binding sites, emphasizing the functional relevance of isolated sites.
Conclusion: Taken together, our in vivo binding and gene expression data support a role for the Su(Hw) protein in maintaining a constant genomic architecture.
Figures
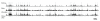





References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

