The Atg5 Atg12 conjugate associates with innate antiviral immune responses
- PMID: 17709747
- PMCID: PMC1955809
- DOI: 10.1073/pnas.0704014104
The Atg5 Atg12 conjugate associates with innate antiviral immune responses
Abstract
Autophagy is an essential process for physiological homeostasis, but its role in viral infection is only beginning to be elucidated. We show here that the Atg5-Atg12 conjugate, a key regulator of the autophagic process, plays an important role in innate antiviral immune responses. Atg5-deficient mouse embryonic fibroblasts (MEFs) were resistant to vesicular stomatitis virus replication, which was largely due to hyperproduction of type I interferons in response to immunostimulatory RNA (isRNA), such as virus-derived, double-stranded, or 5'-phosphorylated RNA. Similar hyperresponse to isRNA was also observed in Atg7-deficient MEFs, in which Atg5-Atg12 conjugation is impaired. Overexpression of Atg5 or Atg12 resulted in Atg5-Atg12 conjugate formation and suppression of isRNA-mediated signaling. Molecular interaction studies indicated that the Atg5-Atg12 conjugate negatively regulates the type I IFN production pathway by direct association with the retinoic acid-inducible gene I (RIG-I) and IFN-beta promoter stimulator 1 (IPS-1) through the caspase recruitment domains (CARDs). Thus, in contrast to its role in promoting the bactericidal process, a component of the autophagic machinery appears to block innate antiviral immune responses, thereby contributing to RNA virus replication in host cells.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
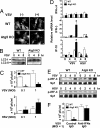




Similar articles
-
The non-canonical role of Atg family members as suppressors of innate antiviral immune signaling.Autophagy. 2008 Jan;4(1):67-9. doi: 10.4161/auto.5055. Epub 2007 Sep 10. Autophagy. 2008. PMID: 17921696
-
Foot-and-mouth disease virus infection suppresses autophagy and NF-кB antiviral responses via degradation of ATG5-ATG12 by 3Cpro.Cell Death Dis. 2017 Jan 19;8(1):e2561. doi: 10.1038/cddis.2016.489. Cell Death Dis. 2017. PMID: 28102839 Free PMC article.
-
Hepatitis B Virus Subverts the Autophagy Elongation Complex Atg5-12/16L1 and Does Not Require Atg8/LC3 Lipidation for Viral Maturation.J Virol. 2018 Mar 14;92(7):e01513-17. doi: 10.1128/JVI.01513-17. Print 2018 Apr 1. J Virol. 2018. PMID: 29367244 Free PMC article.
-
ATG5: A central autophagy regulator implicated in various human diseases.Cell Biochem Funct. 2022 Oct;40(7):650-667. doi: 10.1002/cbf.3740. Epub 2022 Sep 5. Cell Biochem Funct. 2022. PMID: 36062813 Review.
-
Interplay between the cellular autophagy machinery and positive-stranded RNA viruses.Acta Biochim Biophys Sin (Shanghai). 2012 May;44(5):375-84. doi: 10.1093/abbs/gms010. Epub 2012 Feb 16. Acta Biochim Biophys Sin (Shanghai). 2012. PMID: 22343377 Free PMC article. Review.
Cited by
-
Immunologic manifestations of autophagy.J Clin Invest. 2015 Jan;125(1):75-84. doi: 10.1172/JCI73945. Epub 2015 Jan 2. J Clin Invest. 2015. PMID: 25654553 Free PMC article. Review.
-
RNase L induces autophagy via c-Jun N-terminal kinase and double-stranded RNA-dependent protein kinase signaling pathways.J Biol Chem. 2012 Dec 21;287(52):43651-64. doi: 10.1074/jbc.M112.399964. Epub 2012 Oct 29. J Biol Chem. 2012. PMID: 23109342 Free PMC article.
-
MAVS maintains mitochondrial homeostasis via autophagy.Cell Discov. 2016 Aug 16;2:16024. doi: 10.1038/celldisc.2016.24. eCollection 2016. Cell Discov. 2016. PMID: 27551434 Free PMC article.
-
Coxsackievirus A16 elicits incomplete autophagy involving the mTOR and ERK pathways.PLoS One. 2015 Apr 8;10(4):e0122109. doi: 10.1371/journal.pone.0122109. eCollection 2015. PLoS One. 2015. PMID: 25853521 Free PMC article.
-
Autophagy as an innate immune modulator.Immune Netw. 2013 Feb;13(1):1-9. doi: 10.4110/in.2013.13.1.1. Epub 2013 Feb 28. Immune Netw. 2013. PMID: 23559894 Free PMC article.
References
-
- Gutierrez MG, Master SS, Singh SB, Taylor GA, Colombo MI, Deretic V. Cell. 2004;119:753–766. - PubMed
-
- Nakagawa I, Amano A, Mizushima N, Yamamoto A, Yamaguchi H, Kamimoto T, Nara A, Funao J, Nakata M, Tsuda K, et al. Science. 2004;306:1037–1040. - PubMed
-
- Ogawa M, Yoshimori T, Suzuki T, Sagara H, Mizushima N, Sasakawa C. Science. 2005;307:727–731. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

