A new approach to estimate parameters of speciation models with application to apes
- PMID: 17712021
- PMCID: PMC1987350
- DOI: 10.1101/gr.6409707
A new approach to estimate parameters of speciation models with application to apes
Abstract
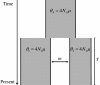
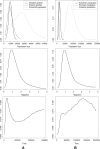
How populations diverge and give rise to distinct species remains a fundamental question in evolutionary biology, with important implications for a wide range of fields, from conservation genetics to human evolution. A promising approach is to estimate parameters of simple speciation models using polymorphism data from multiple loci. Existing methods, however, make a number of assumptions that severely limit their applicability, notably, no gene flow after the populations split and no intralocus recombination. To overcome these limitations, we developed a new Markov chain Monte Carlo method to estimate parameters of an isolation-migration model. The approach uses summaries of polymorphism data at multiple loci surveyed in a pair of diverging populations or closely related species and, importantly, allows for intralocus recombination. To illustrate its potential, we applied it to extensive polymorphism data from populations and species of apes, whose demographic histories are largely unknown. The isolation-migration model appears to provide a reasonable fit to the data. It suggests that the two chimpanzee species became reproductively isolated in allopatry approximately 850 Kya, while Western and Central chimpanzee populations split approximately 440 Kya but continued to exchange migrants. Similarly, Eastern and Western gorillas and Sumatran and Bornean orangutans appear to have experienced gene flow since their splits approximately 90 and over 250 Kya, respectively.
Figures


References
-
- Altschul S., Gish W., Miller W., Meyers E., Lipman D., Gish W., Miller W., Meyers E., Lipman D., Miller W., Meyers E., Lipman D., Meyers E., Lipman D., Lipman D. Basic local alignment search tool. J. Mol. Biol. 1990;215:403–410. - PubMed
-
- Barton N., Bengtsson B., Bengtsson B. The barrier to genetic exchange between hybridising populations. Heredity. 1986;57:357–376. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
