The state of the art in the analysis of two-dimensional gel electrophoresis images
- PMID: 17713763
- PMCID: PMC2279157
- DOI: 10.1007/s00253-007-1128-0
The state of the art in the analysis of two-dimensional gel electrophoresis images
Erratum in
- Appl Microbiol Biotechnol. 2008 May;79(2):329
Abstract
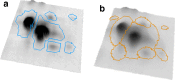
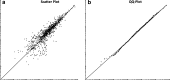
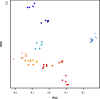
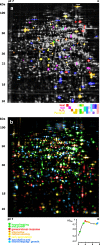
Software-based image analysis is a crucial step in the biological interpretation of two-dimensional gel electrophoresis experiments. Recent significant advances in image processing methods combined with powerful computing hardware have enabled the routine analysis of large experiments. We cover the process starting with the imaging of 2-D gels, quantitation of spots, creation of expression profiles to statistical expression analysis followed by the presentation of results. Challenges for analysis software as well as good practices are highlighted. We emphasize image warping and related methods that are able to overcome the difficulties that are due to varying migration positions of spots between gels. Spot detection, quantitation, normalization, and the creation of expression profiles are described in detail. The recent development of consensus spot patterns and complete expression profiles enables one to take full advantage of statistical methods for expression analysis that are well established for the analysis of DNA microarray experiments. We close with an overview of visualization and presentation methods (proteome maps) and current challenges in the field.
Figures













References
-
- Aittokallio T, Salmi J, Nyman TA, Nevalainen OS. Geometrical distortions in two-dimensional gels: applicable correction method. J Chromatogr B. 2005;815:25–37. - PubMed
-
- Alban A, Davis SO, Bjorkesten L, Andersson C, et al. A novel experimental design for comparative two-dimensional gel analysis: two dimensional difference gel electrophoresis incorporating a pooled internal standard. Proteomics. 2003;3:36–44. - PubMed
-
- Amemiya Y, Miyahara J. Imaging plate illuminates many fields. Nature. 1988;336(6194):89–90. - PubMed
-
- Anderson NL, Taylor J, Scandora AE, Coulter BP, Anderson NG. The TYCHO system for computer analysis of two-dimensional gel electrophoresis patterns. Clin Chem. 1981;27:1807–1820. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

