Specific binding of a Pop6/Pop7 heterodimer to the P3 stem of the yeast RNase MRP and RNase P RNAs
- PMID: 17717080
- PMCID: PMC1986809
- DOI: 10.1261/rna.654407
Specific binding of a Pop6/Pop7 heterodimer to the P3 stem of the yeast RNase MRP and RNase P RNAs
Abstract
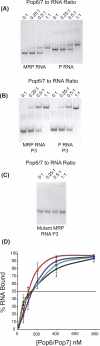
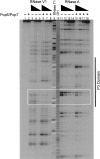
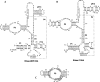
Pop6 and Pop7 are protein subunits of Saccharomyces cerevisiae RNase MRP and RNase P. Here we show that bacterially expressed Pop6 and Pop7 form a soluble heterodimer that binds the RNA components of both RNase MRP and RNase P. Footprint analysis of the interaction between the Pop6/7 heterodimer and the RNase MRP RNA, combined with gel mobility assays, demonstrates that the Pop6/7 complex binds to a conserved region of the P3 domain. Binding of these proteins to the MRP RNA leads to local rearrangement in the structure of the P3 loop and suggests that direct interaction of the Pop6/7 complex with the P3 domain of the RNA components of RNases MRP and P may mediate binding of other protein components. These results suggest a role for a key element in the RNase MRP and RNase P RNAs in protein binding, and demonstrate the feasibility of directly studying RNA-protein interactions in the eukaryotic RNases MRP and P complexes.
Figures

Similar articles
-
Crystallization and preliminary X-ray diffraction analysis of the P3 RNA domain of yeast ribonuclease MRP in a complex with RNase P/MRP protein components Pop6 and Pop7.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2010 Jan 1;66(Pt 1):76-80. doi: 10.1107/S1744309109049707. Epub 2009 Dec 25. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2010. PMID: 20057077 Free PMC article.
-
Active Yeast Telomerase Shares Subunits with Ribonucleoproteins RNase P and RNase MRP.Cell. 2016 May 19;165(5):1171-1181. doi: 10.1016/j.cell.2016.04.018. Epub 2016 May 5. Cell. 2016. PMID: 27156450 Free PMC article.
-
Eukaryotic ribonucleases P/MRP: the crystal structure of the P3 domain.EMBO J. 2010 Feb 17;29(4):761-9. doi: 10.1038/emboj.2009.396. Epub 2010 Jan 14. EMBO J. 2010. PMID: 20075859 Free PMC article.
-
Genetic and biochemical analyses of yeast RNase MRP.Mol Biol Rep. 1995-1996;22(2-3):75-9. doi: 10.1007/BF00988709. Mol Biol Rep. 1995. PMID: 8901491 Review.
-
Structural and functional similarities between MRP and RNase P.Mol Biol Rep. 1995-1996;22(2-3):81-5. doi: 10.1007/BF00988710. Mol Biol Rep. 1995. PMID: 8901492 Review.
Cited by
-
Footprinting analysis of interactions between the largest eukaryotic RNase P/MRP protein Pop1 and RNase P/MRP RNA components.RNA. 2015 Sep;21(9):1591-605. doi: 10.1261/rna.049007.114. Epub 2015 Jul 1. RNA. 2015. PMID: 26135751 Free PMC article.
-
Biogenesis of RNase P RNA from an intron requires co-assembly with cognate protein subunits.Nucleic Acids Res. 2019 Sep 19;47(16):8746-8754. doi: 10.1093/nar/gkz572. Nucleic Acids Res. 2019. PMID: 31287870 Free PMC article.
-
Footprinting analysis demonstrates extensive similarity between eukaryotic RNase P and RNase MRP holoenzymes.RNA. 2008 Aug;14(8):1558-67. doi: 10.1261/rna.1106408. Epub 2008 Jun 25. RNA. 2008. PMID: 18579867 Free PMC article.
-
Crystallization and preliminary X-ray diffraction analysis of the P3 RNA domain of yeast ribonuclease MRP in a complex with RNase P/MRP protein components Pop6 and Pop7.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2010 Jan 1;66(Pt 1):76-80. doi: 10.1107/S1744309109049707. Epub 2009 Dec 25. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2010. PMID: 20057077 Free PMC article.
-
Modular architecture of eukaryotic RNase P and RNase MRP revealed by electron microscopy.Nucleic Acids Res. 2012 Apr;40(7):3275-88. doi: 10.1093/nar/gkr1217. Epub 2011 Dec 13. Nucleic Acids Res. 2012. PMID: 22167472 Free PMC article.
References
-
- Altman, S., Kirsebom, L. Ribonuclease P. In: Gesteland R.F., et al., editors. The RNA world. Cold Spring Harbor Laboratory Press; Cold Spring Harbor, NY: 1999. pp. 351–380.
-
- Forster, A.C., Altman, S. Similar cage-shaped structures for the RNA components of all Ribonuclease P and Ribonuclease MRP enzymes. Cell. 1990;62:407–409. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
