Comparative assessment of plasmid and oligonucleotide DNA substrates in measurement of in vitro base excision repair activity
- PMID: 17720705
- PMCID: PMC2034467
- DOI: 10.1093/nar/gkm639
Comparative assessment of plasmid and oligonucleotide DNA substrates in measurement of in vitro base excision repair activity
Abstract
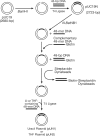
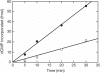
Mammalian base excision repair (BER) is mediated through at least two subpathways designated 'single-nucleotide' (SN) and 'long-patch' (LP) BER (2-nucleotides long/more repair patch). Two forms of DNA substrate are generally used for in vitro BER assays: oligonucleotide- and plasmid-based. For plasmid-based BER assays, the availability of large quantities of substrate DNA with a specific lesion remains the limiting factor. Using sequence-specific endonucleases that cleave only one strand of DNA on a double-stranded DNA substrate, we prepared large quantities of plasmid DNA with a specific lesion. We compared the kinetic features of BER using plasmid and oligonucleotide substrates containing the same lesion and strategic restriction sites around the lesion. The K(m) for plasmid DNA substrate was slightly higher than that for the oligonucleotide substrate, while the V(max) of BER product formation for the plasmid and oligonucleotide substrates was similar. The catalytic efficiency of BER with the oligonucleotide substrate was slightly higher than that with the plasmid substrate. We conclude that there were no significant differences in the catalytic efficiency of in vitro BER measured with plasmid and oligonucleotide substrates. Analysis of the ratio of SN BER to LP BER was addressed using cellular extracts and a novel plasmid substrate.
Figures

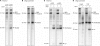



Similar articles
-
Long-patch DNA repair synthesis during base excision repair in mammalian cells.EMBO Rep. 2003 Apr;4(4):363-7. doi: 10.1038/sj.embor.embor796. EMBO Rep. 2003. PMID: 12671676 Free PMC article.
-
Second pathway for completion of human DNA base excision-repair: reconstitution with purified proteins and requirement for DNase IV (FEN1).EMBO J. 1997 Jun 2;16(11):3341-8. doi: 10.1093/emboj/16.11.3341. EMBO J. 1997. PMID: 9214649 Free PMC article.
-
In vitro base excision repair assay using mammalian cell extracts.Methods Mol Biol. 2006;314:377-96. doi: 10.1385/1-59259-973-7:377. Methods Mol Biol. 2006. PMID: 16673895
-
Base damage and single-strand break repair: mechanisms and functional significance of short- and long-patch repair subpathways.DNA Repair (Amst). 2007 Apr 1;6(4):398-409. doi: 10.1016/j.dnarep.2006.10.008. Epub 2006 Nov 28. DNA Repair (Amst). 2007. PMID: 17129767 Review.
-
Roles of base excision repair subpathways in correcting oxidized abasic sites in DNA.FEBS J. 2006 Apr;273(8):1620-9. doi: 10.1111/j.1742-4658.2006.05192.x. FEBS J. 2006. PMID: 16623699 Review.
Cited by
-
FEN1 functions in long patch base excision repair under conditions of oxidative stress in vertebrate cells.Mol Cancer Res. 2010 Feb;8(2):204-15. doi: 10.1158/1541-7786.MCR-09-0253. Epub 2010 Feb 9. Mol Cancer Res. 2010. PMID: 20145043 Free PMC article.
-
Is the oxidative DNA damage level of human lymphocyte correlated with the antioxidant capacity of serum or the base excision repair activity of lymphocyte?Oxid Med Cell Longev. 2013;2013:237583. doi: 10.1155/2013/237583. Epub 2013 Nov 14. Oxid Med Cell Longev. 2013. PMID: 24349611 Free PMC article.
-
Oxidative stress alters base excision repair pathway and increases apoptotic response in apurinic/apyrimidinic endonuclease 1/redox factor-1 haploinsufficient mice.Free Radic Biol Med. 2009 Jun 1;46(11):1488-99. doi: 10.1016/j.freeradbiomed.2009.02.021. Epub 2009 Mar 3. Free Radic Biol Med. 2009. PMID: 19268524 Free PMC article.
-
Automated AFM analysis of DNA bending reveals initial lesion sensing strategies of DNA glycosylases.Sci Rep. 2020 Sep 23;10(1):15484. doi: 10.1038/s41598-020-72102-7. Sci Rep. 2020. PMID: 32968112 Free PMC article.
-
Nucleosomes determine their own patch size in base excision repair.Sci Rep. 2016 Jun 6;6:27122. doi: 10.1038/srep27122. Sci Rep. 2016. PMID: 27265863 Free PMC article.
References
-
- Lindahl T. DNA repair enzymes. Annu. Rev. Biochem. 1982;51:61–87. - PubMed
-
- Lindahl T, Wood RD. Quality control by DNA repair. Science. 1999;286:1897–1905. - PubMed
-
- Biade S, Sobol RW, Wilson SH, Matsumoto Y. Impairment of proliferating cell nuclear antigen-dependent apurinic/apyrimidinic site repair on linear DNA. J. Biol. Chem. 1998;273:898–902. - PubMed
-
- Fortini P, Pascucci B, Parlanti E, Sobol RW, Wilson SH, Dogliotti E. Different DNA polymerases are involved in the short- and long-patch base excision repair in mammalian cells. Biochemistry. 1998;37:3575–3580. - PubMed

