Activation of yeaR-yoaG operon transcription by the nitrate-responsive regulator NarL is independent of oxygen- responsive regulator Fnr in Escherichia coli K-12
- PMID: 17720788
- PMCID: PMC2168752
- DOI: 10.1128/JB.00953-07
Activation of yeaR-yoaG operon transcription by the nitrate-responsive regulator NarL is independent of oxygen- responsive regulator Fnr in Escherichia coli K-12
Abstract
The facultative aerobe Escherichia coli K-12 can use respiratory nitrate ammonification to generate energy during anaerobic growth. The toxic compound nitric oxide is a by-product of this metabolism. Previous transcript microarray studies identified the yeaR-yoaG operon, encoding proteins of unknown function, among genes whose transcription is induced in response to nitrate, nitrite, or nitric oxide. Nitrate and nitrite regulate anaerobic respiratory gene expression through the NarX-NarL and NarQ-NarP two-component systems. All known Nar-activated genes also require the oxygen-responsive Fnr transcription activator. However, previous studies indicated that yeaR-yoaG operon transcription does not require Fnr activation. Here, we report results from mutational analyses demonstrating that yeaR-yoaG operon transcription is activated by phospho-NarL protein independent of the Fnr protein. The phospho-NarL protein binding site is centered at position -43.5 with respect to the transcription initiation site. Expression from the Shewanella oneidensis MR-1 nnrS gene promoter, cloned into E. coli, similarly was activated by phospho-NarL protein independent of the Fnr protein. Recently, yeaR-yoaG operon transcription was shown to be regulated by the nitric oxide-responsive NsrR repressor (N. Filenko et al., J. Bacteriol. 189:4410-4417, 2007). Our mutational analyses reveal the individual contributions of the Nar and NsrR regulators to overall yeaR-yoaG operon expression and document the NsrR operator centered at position -32. Thus, control of yeaR-yoaG operon transcription provides an example of overlapping regulation by nitrate and nitrite, acting through the Nar regulatory system, and nitric oxide, acting through the NsrR repressor.
Figures

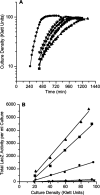
References
-
- Barnard, A., A. Wolfe, and S. Busby. 2004. Regulation at complex bacterial promoters: how bacteria use different promoter organizations to produce different regulatory outcomes. Curr. Opin. Microbiol. 7:102-108. - PubMed
-
- Bartnikas, T. B., Y. Wang, T. Bobo, A. Veselov, C. P. Scholes, and J. P. Shapleigh. 2002. Characterization of a member of the NnrR regulon in Rhodobacter sphaeroides 2.4.3 encoding a haem-copper protein. Microbiology 148:825-833. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

