Cryptosporidium genotypes in wildlife from a new york watershed
- PMID: 17720824
- PMCID: PMC2075072
- DOI: 10.1128/AEM.01034-07
Cryptosporidium genotypes in wildlife from a new york watershed
Abstract
To identify the animal sources for Cryptosporidium contamination, we genotyped Cryptosporidium spp. in wildlife from the watershed of the New York City drinking water supply, using a small-subunit rRNA gene-based PCR-restriction fragment length polymorphism analysis and DNA sequencing. A total of 541 specimens from 38 species of wildlife were analyzed. One hundred and eleven (20.5%) of the wildlife specimens were PCR positive. Altogether, 21 Cryptosporidium genotypes were found in wildlife samples, 11 of which were previously found in storm runoff in the watershed, and six of these 11 were from storm water genotypes of unknown animal origin. Four new genotypes were found, and the animal hosts for four storm water genotypes were expanded. With the exception of the cervine genotype, most genotypes were found in a limited number of animal species and have no major public health significance.
Figures

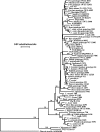
References
-
- Atwill, E. R., M. D. Pereira, L. H. Alonso, C. Elmi, W. B. Epperson, R. Smith, W. Riggs, L. V. Carpenter, D. A. Dargatz, and B. Hoar. 2006. Environmental load of Cryptosporidium parvum oocysts from cattle manure in feedlots from the central and western United States. J. Environ. Qual. 35:200-206. - PubMed
-
- Bajer, A., S. Caccio, M. Bednarska, J. M. Behnke, N. J. Pieniazek, and E. Sinski. 2003. Preliminary molecular characterization of Cryptosporidium parvum isolates of wildlife rodents from Poland. J. Parasitol. 89:1053-1055. - PubMed
-
- Bednarska, M., A. Bajer, K. Kulis, and E. Sinski. 2003. Biological characterisation of Cryptosporidium parvum isolates of wildlife rodents in Poland. Ann. Agric. Environ. Med. 10:163-169. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

