Large-scale, lineage-specific expansion of a bric-a-brac/tramtrack/broad complex ubiquitin-ligase gene family in rice
- PMID: 17720868
- PMCID: PMC2002615
- DOI: 10.1105/tpc.107.051300
Large-scale, lineage-specific expansion of a bric-a-brac/tramtrack/broad complex ubiquitin-ligase gene family in rice
Abstract
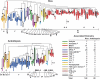
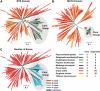
Selective ubiquitination of proteins is directed by diverse families of ubiquitin-protein ligases (or E3s) in plants. One important type uses Cullin-3 as a scaffold to assemble multisubunit E3 complexes containing one of a multitude of bric-a-brac/tramtrack/broad complex (BTB) proteins that function as substrate recognition factors. We previously described the 80-member BTB gene superfamily in Arabidopsis thaliana. Here, we describe the complete BTB superfamily in rice (Oryza sativa spp japonica cv Nipponbare) that contains 149 BTB domain-encoding genes and 43 putative pseudogenes. Amino acid sequence comparisons of the rice and Arabidopsis superfamilies revealed a near equal repertoire of putative substrate recognition module types. However, phylogenetic comparisons detected numerous gene duplication and/or loss events since the rice and Arabidopsis BTB lineages split, suggesting possible functional specialization within individual BTB families. In particular, a major expansion and diversification of a subset of BTB proteins containing Meprin and TRAF homology (MATH) substrate recognition sites was evident in rice and other monocots that likely occurred following the monocot/dicot split. The MATH domain of a subset appears to have evolved significantly faster than those in a smaller core subset that predates flowering plants, suggesting that the substrate recognition module in many monocot MATH-BTB E3s are diversifying to ubiquitinate a set of substrates that are themselves rapidly changing. Intriguing possibilities include pathogen proteins attempting to avoid inactivation by the monocot host.
Figures






References
-
- Aguilar, R.C., and Wendland, B. (2003). Ubiquitin: Not just for proteasomes anymore. Curr. Opin. Cell Biol. 15 184–190. - PubMed
-
- Angot, A., Peeters, N., Lechner, E., Vailleau, F., Baud, C., Gentzbittel, L., Sartorel, E., Genschik, P., Boucher, C., and Genin, S. (2006). Ralstonia solanacearum requires F-box-like domain-containing type III effectors to promote disease on several host plants. Proc. Natl. Acad. Sci. USA 103 14620–14625. - PMC - PubMed
-
- Aravind, L., and Koonin, E.V. (1999). Fold prediction and evolutionary analysis of the POZ domain: Structural and evolutionary relationship with the potassium channel tetramerization domain. J. Mol. Biol. 285 1353–1361. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

