Genetic and haplotypic structure in 14 European and African cattle breeds
- PMID: 17720924
- PMCID: PMC2034613
- DOI: 10.1534/genetics.107.075804
Genetic and haplotypic structure in 14 European and African cattle breeds
Abstract
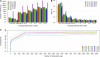
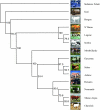
To evaluate and compare the extent of LD in cattle, 1536 SNPs, mostly localized on BTA03, were detected in silico from available sequence data using two different methods and genotyped on samples from 14 distinct breeds originating from Europe and Africa. Only 696 SNPs could be validated, confirming the importance of trace-quality information for the in silico detection. Most of the validated SNPs were informative in several breeds and were used for a detailed description of their genetic structure and relationships. Results obtained were in agreement with previous studies performed on microsatellite markers and using larger samples. In addition, the majority of the validated SNPs could be mapped precisely, reaching an average density of one marker every 311 kb. This allowed us to analyze the extent of LD in the different breeds. Decrease of LD with physical distance across breeds revealed footprints of ancestral LD at short distances (<10 kb). As suggested by the haplotype block structure, these ancestral blocks are organized, within a breed, into larger blocks of a few hundred kilobases. In practice, such a structure similar to that already reported in dogs makes it possible to develop a chip of <300,000 SNPs, which should be efficient for mapping purposes in most cattle breeds.
Figures
Similar articles
-
High-resolution haplotype block structure in the cattle genome.BMC Genet. 2009 Apr 24;10:19. doi: 10.1186/1471-2156-10-19. BMC Genet. 2009. PMID: 19393054 Free PMC article.
-
The extent of linkage disequilibrium in beef cattle breeds using high-density SNP genotypes.Genet Sel Evol. 2014 Mar 24;46(1):22. doi: 10.1186/1297-9686-46-22. Genet Sel Evol. 2014. PMID: 24661366 Free PMC article.
-
Linkage disequilibrium decay and haplotype block structure in the pig.Genetics. 2008 May;179(1):569-79. doi: 10.1534/genetics.107.084277. Genetics. 2008. PMID: 18493072 Free PMC article.
-
High density linkage disequilibrium maps of chromosome 14 in Holstein and Angus cattle.BMC Genet. 2008 Jul 8;9:45. doi: 10.1186/1471-2156-9-45. BMC Genet. 2008. PMID: 18611270 Free PMC article.
-
Linkage disequilibrium and haplotype block structure in a composite beef cattle breed.BMC Genomics. 2014;15 Suppl 7(Suppl 7):S6. doi: 10.1186/1471-2164-15-S7-S6. Epub 2014 Oct 27. BMC Genomics. 2014. PMID: 25573652 Free PMC article.
Cited by
-
Whole genome detection of recent selection signatures in Sarabi cattle: a unique Iranian taurine breed.Genes Genomics. 2020 Feb;42(2):203-215. doi: 10.1007/s13258-019-00888-6. Epub 2019 Dec 5. Genes Genomics. 2020. PMID: 31808064
-
The genetic history of Mayotte and Madagascar cattle breeds mirrors the complex pattern of human exchanges in Western Indian Ocean.G3 (Bethesda). 2022 Apr 4;12(4):jkac029. doi: 10.1093/g3journal/jkac029. G3 (Bethesda). 2022. PMID: 35137043 Free PMC article.
-
Mapping of a milk production quantitative trait locus to a 1.056 Mb region on bovine chromosome 5 in the Fleckvieh dual purpose cattle breed.Genet Sel Evol. 2011 Feb 24;43(1):8. doi: 10.1186/1297-9686-43-8. Genet Sel Evol. 2011. PMID: 21349166 Free PMC article.
-
Assessing signatures of selection through variation in linkage disequilibrium between taurine and indicine cattle.Genet Sel Evol. 2014 Mar 4;46(1):19. doi: 10.1186/1297-9686-46-19. Genet Sel Evol. 2014. PMID: 24592996 Free PMC article.
-
Linkage Disequilibrium and Effective Population Size of Buffalo Populations of Iran, Turkey, Pakistan, and Egypt Using a Medium Density SNP Array.Front Genet. 2021 Dec 7;12:608186. doi: 10.3389/fgene.2021.608186. eCollection 2021. Front Genet. 2021. PMID: 34950186 Free PMC article.
References
-
- Andersson, L., and M. Georges, 2004. Domestic-animal genomics: deciphering the genetics of complex traits. Nat. Rev. Genet. 5: 202–212. - PubMed
-
- Ardlie, K. G., L. Kruglyak and M. Seielstad, 2002. Patterns of linkage disequilibrium in the human genome. Nat. Rev. Genet. 3: 299–309. - PubMed
-
- Barrett, J. C., B. Fry, J. Maller and M. J. Daly, 2005. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21: 263–265. - PubMed
-
- Belkhir, K., P. Borsa, L. Chikhi, N. Raufaste and F. Bonhomme, 2004. GENETIX, logiciel sous WindowsTM pour la génétique des populations. Université de Montpellier II, Montpellier, France.
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

