Global gene expression analysis of the shoot apical meristem of maize (Zea mays L.)
- PMID: 17764504
- PMCID: PMC2156186
- DOI: 10.1111/j.1365-313X.2007.03244.x
Global gene expression analysis of the shoot apical meristem of maize (Zea mays L.)
Abstract
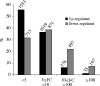
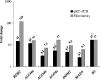
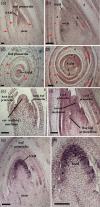
All above-ground plant organs are derived from shoot apical meristems (SAMs). Global analyses of gene expression were conducted on maize (Zea mays L.) SAMs to identify genes preferentially expressed in the SAM. The SAMs were collected from 14-day-old B73 seedlings via laser capture microdissection (LCM). The RNA samples extracted from LCM-collected SAMs and from seedlings were hybridized to microarrays spotted with 37 660 maize cDNAs. Approximately 30% (10 816) of these cDNAs were prepared as part of this study from manually dissected B73 maize apices. Over 5000 expressed sequence tags (ESTs) (about 13% of the total) were differentially expressed (P < 0.0001) between SAMs and seedlings. Of these, 2783 and 2248 ESTs were up- and down-regulated in the SAM, respectively. The expression in the SAM of several of the differentially expressed ESTs was validated via quantitative RT-PCR and/or in situ hybridization. The up-regulated ESTs included many regulatory genes including transcription factors, chromatin remodeling factors and components of the gene-silencing machinery, as well as about 900 genes with unknown functions. Surprisingly, transcripts that hybridized to 62 retrotransposon-related cDNAs were also substantially up-regulated in the SAM. Complementary DNAs derived from the LCM-collected SAMs were sequenced to identify additional genes that are expressed in the SAM. This generated around 550 000 ESTs (454-SAM ESTs) from two genotypes. Consistent with the microarray results, approximately 14% of the 454-SAM ESTs from B73 were retrotransposon-related. Possible roles of genes that are preferentially expressed in the SAM are discussed.
Figures

Similar articles
-
Genome-wide analysis of gene expression in soybean shoot apical meristem.Plant Mol Biol. 2009 Apr;69(6):711-27. doi: 10.1007/s11103-008-9450-1. Epub 2008 Dec 30. Plant Mol Biol. 2009. PMID: 19115044
-
Gene discovery and annotation using LCM-454 transcriptome sequencing.Genome Res. 2007 Jan;17(1):69-73. doi: 10.1101/gr.5145806. Epub 2006 Nov 9. Genome Res. 2007. PMID: 17095711 Free PMC article.
-
Ontogeny of the maize shoot apical meristem.Plant Cell. 2012 Aug;24(8):3219-34. doi: 10.1105/tpc.112.099614. Epub 2012 Aug 21. Plant Cell. 2012. PMID: 22911570 Free PMC article.
-
Decoding maize meristems maintenance and differentiation: integrating single-cell and spatial omics.J Genet Genomics. 2025 Mar;52(3):319-333. doi: 10.1016/j.jgg.2025.01.012. Epub 2025 Feb 5. J Genet Genomics. 2025. PMID: 39921079 Review.
-
All together now, a magical mystery tour of the maize shoot meristem.Curr Opin Plant Biol. 2018 Oct;45(Pt A):26-35. doi: 10.1016/j.pbi.2018.04.010. Epub 2018 May 17. Curr Opin Plant Biol. 2018. PMID: 29778985 Review.
Cited by
-
Characterization of WRKY co-regulatory networks in rice and Arabidopsis.BMC Plant Biol. 2009 Sep 22;9:120. doi: 10.1186/1471-2229-9-120. BMC Plant Biol. 2009. PMID: 19772648 Free PMC article.
-
Genomic convergence analysis of schizophrenia: mRNA sequencing reveals altered synaptic vesicular transport in post-mortem cerebellum.PLoS One. 2008;3(11):e3625. doi: 10.1371/journal.pone.0003625. Epub 2008 Nov 5. PLoS One. 2008. PMID: 18985160 Free PMC article.
-
Floral primordia-targeted ACS (1-aminocyclopropane-1-carboxylate synthase) expression in transgenic Cucumis melo implicates fine tuning of ethylene production mediating unisexual flower development.Planta. 2014 Oct;240(4):797-808. doi: 10.1007/s00425-014-2118-y. Epub 2014 Jul 29. Planta. 2014. PMID: 25066672
-
RNASeq Analysis of the Shoot Apex of Flax (Linum usitatissimum) to Identify Phloem Fiber Specification Genes.Front Plant Sci. 2016 Jun 28;7:950. doi: 10.3389/fpls.2016.00950. eCollection 2016. Front Plant Sci. 2016. PMID: 27446177 Free PMC article. No abstract available.
-
The naked endosperm genes encode duplicate INDETERMINATE domain transcription factors required for maize endosperm cell patterning and differentiation.Plant Physiol. 2015 Feb;167(2):443-56. doi: 10.1104/pp.114.251413. Epub 2014 Dec 31. Plant Physiol. 2015. PMID: 25552497 Free PMC article.
References
-
- Abbe EC. The growth of the shoot apex in maize: embryogeny. Am. J. Bot. 1954;41:285–293.
-
- Alleman M. An RNA-dependent RNA polymerase is required for paramutation in maize. Nature. 2006;442:295–298. - PubMed
-
- Alvarez-Buylla ER. MADS-box gene evolution beyond flowers: expression in pollen, endosperm, guard cells, roots and trichomes. Plant J. 2000;24:457–466. - PubMed
-
- Bäurle I. Apical meristems: the plant’s fountain of youth. BioEssays. 2003;25:961–970. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

