The mitochondrial genome of Phallusia mammillata and Phallusia fumigata (Tunicata, Ascidiacea): high genome plasticity at intra-genus level
- PMID: 17764550
- PMCID: PMC2220002
- DOI: 10.1186/1471-2148-7-155
The mitochondrial genome of Phallusia mammillata and Phallusia fumigata (Tunicata, Ascidiacea): high genome plasticity at intra-genus level
Abstract
Background: Within Chordata, the subphyla Vertebrata and Cephalochordata (lancelets) are characterized by a remarkable stability of the mitochondrial (mt) genome, with constancy of gene content and almost invariant gene order, whereas the limited mitochondrial data on the subphylum Tunicata suggest frequent and extensive gene rearrangements, observed also within ascidians of the same genus.
Results: To confirm this evolutionary trend and to better understand the evolutionary dynamics of the mitochondrial genome in Tunicata Ascidiacea, we have sequenced and characterized the complete mt genome of two congeneric ascidian species, Phallusia mammillata and Phallusia fumigata (Phlebobranchiata, Ascidiidae). The two mtDNAs are surprisingly rearranged, both with respect to one another and relative to those of other tunicates and chordates, with gene rearrangements affecting both protein-coding and tRNA genes. The new data highlight the extraordinary variability of ascidian mt genome in base composition, tRNA secondary structure, tRNA gene content, and non-coding regions (number, size, sequence and location). Indeed, both Phallusia genomes lack the trnD gene, show loss/acquisition of DHU-arm in two tRNAs, and have a G+C content two-fold higher than other ascidians. Moreover, the mt genome of P. fumigata presents two identical copies of trnI, an extra tRNA gene with uncertain amino acid specificity, and four almost identical sequence regions. In addition, a truncated cytochrome b, lacking a C-terminal tail that commonly protrudes into the mt matrix, has been identified as a new mt feature probably shared by all tunicates.
Conclusion: The frequent occurrence of major gene order rearrangements in ascidians both at high taxonomic level and within the same genus makes this taxon an excellent model to study the mechanisms of gene rearrangement, and renders the mt genome an invaluable phylogenetic marker to investigate molecular biodiversity and speciation events in this largely unexplored group of basal chordates.
Figures



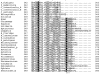

References
-
- Serb JM, Lydeard C. Complete mtDNA sequence of the North American freshwater mussel, Lampsilis ornata (Unionidae): an examination of the evolution and phylogenetic utility of mitochondrial genome organization in Bivalvia (Mollusca) Mol Biol Evol. 2003;20:1854–1866. doi: 10.1093/molbev/msg218. - DOI - PubMed

