Antigen 84, an effector of pleiomorphism in Mycobacterium smegmatis
- PMID: 17766411
- PMCID: PMC2168712
- DOI: 10.1128/JB.00726-07
Antigen 84, an effector of pleiomorphism in Mycobacterium smegmatis
Abstract
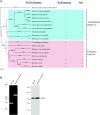
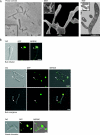
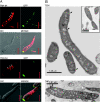
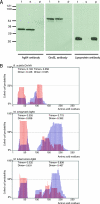
While in most rod-shaped bacteria, morphology is based on MreB-like proteins that form an actin-like cytoskeletal scaffold for cell wall biosynthesis, the factors that determine the more flexible rod-like shape in actinobacteria such as Mycobacterium species are unknown. Here we show that a Mycobacterium smegmatis protein homologous to eubacterial DivIVA-like proteins, including M. tuberculosis antigen 84 (Ag84), localized symmetrically to centers of peptidoglycan biosynthesis at the poles and septa. Controlled gene disruption experiments indicated that the gene encoding Ag84, wag31, was essential; when overexpressed, cells became longer and wider, with Ag84 asymmetrically distributed at one pole. Many became grossly enlarged, bowling-pin-shaped cells having up to 80-fold-increased volume. In these cells, Ag84 accumulated predominantly at a bulbous pole that was apparently generated by uncontrolled cell wall expansion. In some cells, Ag84 was associated with exceptional sites of cell wall expansion (buds) that evolved into branches. M. bovis BCG Ag84 was able to form oligomers in vitro, perhaps reflecting its superstructure in vivo. These data suggested a role for Ag84 in cell division and modulating cell shape in pleiomorphic actinobacteria.
Figures





Similar articles
-
The Wag31 protein interacts with AccA3 and coordinates cell wall lipid permeability and lipophilic drug resistance in Mycobacterium smegmatis.Biochem Biophys Res Commun. 2014 Jun 6;448(3):255-60. doi: 10.1016/j.bbrc.2014.04.116. Epub 2014 Apr 30. Biochem Biophys Res Commun. 2014. PMID: 24792177
-
Mycobacterium tuberculosis Rv1302 and Mycobacterium smegmatis MSMEG_4947 have WecA function and MSMEG_4947 is required for the growth of M. smegmatis.FEMS Microbiol Lett. 2010 Sep 1;310(1):54-61. doi: 10.1111/j.1574-6968.2010.02045.x. Epub 2010 Jun 23. FEMS Microbiol Lett. 2010. PMID: 20637039
-
Rv1818c-encoded PE_PGRS protein of Mycobacterium tuberculosis is surface exposed and influences bacterial cell structure.Mol Microbiol. 2004 May;52(3):725-33. doi: 10.1111/j.1365-2958.2004.04007.x. Mol Microbiol. 2004. PMID: 15101979
-
Comparison of mammalian cell entry operons of mycobacteria: in silico analysis and expression profiling.FEMS Immunol Med Microbiol. 2005 Feb 1;43(2):185-95. doi: 10.1016/j.femsim.2004.08.013. FEMS Immunol Med Microbiol. 2005. PMID: 15681149
-
Is Longitudinal Division in Rod-Shaped Bacteria a Matter of Swapping Axis?Front Microbiol. 2018 May 8;9:822. doi: 10.3389/fmicb.2018.00822. eCollection 2018. Front Microbiol. 2018. PMID: 29867786 Free PMC article. Review.
Cited by
-
Phosphorylation of a novel cytoskeletal protein (RsmP) regulates rod-shaped morphology in Corynebacterium glutamicum.J Biol Chem. 2010 Sep 17;285(38):29387-97. doi: 10.1074/jbc.M110.154427. Epub 2010 Jul 9. J Biol Chem. 2010. PMID: 20622015 Free PMC article.
-
A cell wall synthase accelerates plasma membrane partitioning in mycobacteria.Elife. 2023 Sep 4;12:e81924. doi: 10.7554/eLife.81924. Elife. 2023. PMID: 37665120 Free PMC article.
-
EF1025, a Hypothetical Protein From Enterococcus faecalis, Interacts With DivIVA and Affects Cell Length and Cell Shape.Front Microbiol. 2020 Feb 12;11:83. doi: 10.3389/fmicb.2020.00083. eCollection 2020. Front Microbiol. 2020. PMID: 32117116 Free PMC article.
-
Polar growth in the Alphaproteobacterial order Rhizobiales.Proc Natl Acad Sci U S A. 2012 Jan 31;109(5):1697-701. doi: 10.1073/pnas.1114476109. Epub 2012 Jan 17. Proc Natl Acad Sci U S A. 2012. PMID: 22307633 Free PMC article.
-
Bacterial conversion of folinic acid is required for antifolate resistance.J Biol Chem. 2011 Apr 29;286(17):15377-90. doi: 10.1074/jbc.M111.231076. Epub 2011 Mar 3. J Biol Chem. 2011. PMID: 21372133 Free PMC article.
References
-
- Balasubramanian, M. K., D. M. Helfman, and S. M. Hemmingsen. 1992. A new tropomyosin essential for cytokinesis in the fission yeast S. pombe. Nature 360:84-87. - PubMed
-
- Bardarov, S., S. Bardarov, Jr., M. S. Pavelka, Jr., V. Sambandamurthy, M. Larsen, J. Tufariello, J. Chan, G. Hatfull, and W. R. Jacobs, Jr. 2002. Specialized transduction: an efficient method for generating marked and unmarked targeted gene disruptions in Mycobacterium tuberculosis, M. bovis BCG and M. smegmatis. Microbiology 148:3007-3017. - PubMed
-
- Burnette, W. N. 1981. “Western blotting”: electrophoretic transfer of proteins from sodium dodecyl sulfate-polyacrylamide gels to unmodified nitrocellulose and radiographic detection with antibody and radioiodinated protein A. Anal. Biochem. 112:195-203. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

