Identification of novel surface proteins of Anaplasma phagocytophilum by affinity purification and proteomics
- PMID: 17766422
- PMCID: PMC2168727
- DOI: 10.1128/JB.00866-07
Identification of novel surface proteins of Anaplasma phagocytophilum by affinity purification and proteomics
Abstract
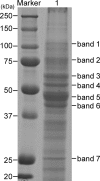
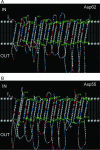
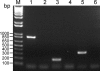
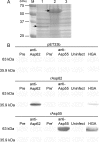
Anaplasma phagocytophilum is the etiologic agent of human granulocytic anaplasmosis (HGA), one of the major tick-borne zoonoses in the United States. The surface of A. phagocytophilum plays a crucial role in subverting the hostile host cell environment. However, except for the P44/Msp2 outer membrane protein family, the surface components of A. phagocytophilum are largely unknown. To identify the major surface proteins of A. phagocytophilum, a membrane-impermeable, cleavable biotin reagent, sulfosuccinimidyl-2-[biotinamido]ethyl-1,3-dithiopropionate (Sulfo-NHS-SS-Biotin), was used to label intact bacteria. The biotinylated bacterial surface proteins were isolated by streptavidin agarose affinity purification and then separated by electrophoresis, followed by capillary liquid chromatography-nanospray tandem mass spectrometry analysis. Among the major proteins captured by affinity purification were five A. phagocytophilum proteins, Omp85, hypothetical proteins APH_0404 (designated Asp62) and APH_0405 (designated Asp55), P44 family proteins, and Omp-1A. The surface exposure of Asp62 and Asp55 was verified by immunofluorescence microscopy. Recombinant Asp62 and Asp55 proteins were recognized by an HGA patient serum. Anti-Asp62 and anti-Asp55 peptide sera partially neutralized A. phagocytophilum infection of HL-60 cells in vitro. We found that the Asp62 and Asp55 genes were cotranscribed and conserved among members of the family Anaplasmataceae. With the exception of P44-18, all of the proteins were newly revealed major surface-exposed proteins whose study should facilitate understanding the interaction between A. phagocytophilum and the host. These proteins may serve as targets for development of chemotherapy, diagnostics, and vaccines.
Figures


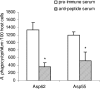
References
-
- Alleman, A. R., A. F. Barbet, H. L. Sorenson, N. I. Strik, H. L. Wamsley, S. J. Wong, R. Chandrashaker, F. P. Gaschen, N. Luckshander, and A. Bjoersdorff. 2006. Cloning and expression of the gene encoding the major surface protein 5 (MSP5) of Anaplasma phagocytophilum and potential application for serodiagnosis. Vet. Clin. Pathol. 35:418-425. - PubMed
-
- Baron, C. 2006. VirB8: a conserved type IV secretion system assembly factor and drug target. Biochem. Cell Biol. 84:890-899. - PubMed
-
- Bendtsen, J. D., H. Nielsen, G. von Heijne, and S. Brunak. 2004. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 340:783-795. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

