Heritable stochastic switching revealed by single-cell genealogy
- PMID: 17803359
- PMCID: PMC1964776
- DOI: 10.1371/journal.pbio.0050239
Heritable stochastic switching revealed by single-cell genealogy
Abstract
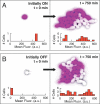
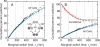
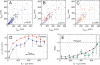
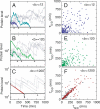
The partitioning and subsequent inheritance of cellular factors like proteins and RNAs is a ubiquitous feature of cell division. However, direct quantitative measures of how such nongenetic inheritance affects subsequent changes in gene expression have been lacking. We tracked families of the yeast Saccharomyces cerevisiae as they switch between two semi-stable epigenetic states. We found that long after two cells have divided, they continued to switch in a synchronized manner, whereas individual cells have exponentially distributed switching times. By comparing these results to a Poisson process, we show that the time evolution of an epigenetic state depends initially on inherited factors, with stochastic processes requiring several generations to decorrelate closely related cells. Finally, a simple stochastic model demonstrates that a single fluctuating regulatory protein that is synthesized in large bursts can explain the bulk of our results.
Conflict of interest statement
Figures

Similar articles
-
Adaptive Potential of Epigenetic Switching During Adaptation to Fluctuating Environments.Genome Biol Evol. 2022 May 3;14(5):evac065. doi: 10.1093/gbe/evac065. Genome Biol Evol. 2022. PMID: 35567483 Free PMC article.
-
Stochastic switching as a survival strategy in fluctuating environments.Nat Genet. 2008 Apr;40(4):471-5. doi: 10.1038/ng.110. Epub 2008 Mar 23. Nat Genet. 2008. PMID: 18362885
-
Enhancement of cellular memory by reducing stochastic transitions.Nature. 2005 May 12;435(7039):228-32. doi: 10.1038/nature03524. Nature. 2005. PMID: 15889097
-
Phenotypic heterogeneity: differential stress resistance among individual cells of the yeast Saccharomyces cerevisiae.Microbiology (Reading). 2002 Feb;148(Pt 2):345-351. doi: 10.1099/00221287-148-2-345. Microbiology (Reading). 2002. PMID: 11832498 Review. No abstract available.
-
Sharing the cell's bounty - organelle inheritance in yeast.J Cell Sci. 2015 Feb 15;128(4):621-30. doi: 10.1242/jcs.151423. Epub 2015 Jan 22. J Cell Sci. 2015. PMID: 25616900 Review.
Cited by
-
Bridging time scales in cellular decision making with a stochastic bistable switch.BMC Syst Biol. 2010 Aug 9;4:108. doi: 10.1186/1752-0509-4-108. BMC Syst Biol. 2010. PMID: 20696063 Free PMC article.
-
Hybrid stochastic simplifications for multiscale gene networks.BMC Syst Biol. 2009 Sep 7;3:89. doi: 10.1186/1752-0509-3-89. BMC Syst Biol. 2009. PMID: 19735554 Free PMC article.
-
Kinetics of genetic switching into the state of bacterial competence.Biophys J. 2009 Feb;96(3):1178-88. doi: 10.1016/j.bpj.2008.10.034. Biophys J. 2009. PMID: 19186153 Free PMC article.
-
Survival of Phenotypic Information during Cellular Growth Transitions.ACS Synth Biol. 2016 Aug 19;5(8):810-6. doi: 10.1021/acssynbio.5b00229. Epub 2016 Mar 7. ACS Synth Biol. 2016. PMID: 26910476 Free PMC article.
-
Stochastic modeling of biochemical systems with multistep reactions using state-dependent time delay.Sci Rep. 2016 Aug 24;6:31909. doi: 10.1038/srep31909. Sci Rep. 2016. PMID: 27553753 Free PMC article.
References
-
- Ray DT, McCreight JD. Yellow-tip: a cytoplasmic inherited trait in melon (Cucumis melo L.) J Hered. 1996;87:245–247.
-
- Gottschling DE, Aparicio OM, Billington BL, Zakian VA. Position effect at S. cerevisiae telomeres: Reversible repression of Pol II transcription. Cell. 1990;63:751–762. - PubMed
-
- Patino MM, Liu JJ, Glover JR, Lindquist S. Support for the prion hypothesis for inheritance of a phenotypic trait in yeast. Science. 1996;273:622–626. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

