Interaction of decay-accelerating factor with coxsackievirus B3
- PMID: 17804498
- PMCID: PMC2169128
- DOI: 10.1128/JVI.00931-07
Interaction of decay-accelerating factor with coxsackievirus B3
Abstract
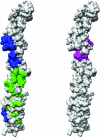
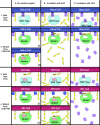
Many entero-, parecho-, and rhinoviruses use immunoglobulin (Ig)-like receptors that bind into the viral canyon and are required to initiate viral uncoating during infection. However, some of these viruses use an alternative or additional receptor that binds outside the canyon. Both the coxsackievirus-adenovirus receptor (CAR), an Ig-like molecule that binds into the viral canyon, and decay-accelerating factor (DAF) have been identified as cellular receptors for coxsackievirus B3 (CVB3). A cryoelectron microscopy reconstruction of a variant of CVB3 complexed with DAF shows full occupancy of the DAF receptor in each of 60 binding sites. The DAF molecule bridges the canyon, blocking the CAR binding site and causing the two receptors to compete with one another. The binding site of DAF on CVB3 differs from the binding site of DAF on the surface of echoviruses, suggesting independent evolutionary processes.
Figures






Similar articles
-
The crystal structure of a coxsackievirus B3-RD variant and a refined 9-angstrom cryo-electron microscopy reconstruction of the virus complexed with decay-accelerating factor (DAF) provide a new footprint of DAF on the virus surface.J Virol. 2012 Dec;86(23):12571-81. doi: 10.1128/JVI.01592-12. Epub 2012 Sep 12. J Virol. 2012. PMID: 22973031 Free PMC article.
-
Interaction of decay-accelerating factor with echovirus 7.J Virol. 2010 Dec;84(24):12665-74. doi: 10.1128/JVI.00837-10. Epub 2010 Sep 29. J Virol. 2010. PMID: 20881044 Free PMC article.
-
Structure of decay-accelerating factor bound to echovirus 7: a virus-receptor complex.Proc Natl Acad Sci U S A. 2002 Aug 6;99(16):10325-9. doi: 10.1073/pnas.152161599. Epub 2002 Jul 15. Proc Natl Acad Sci U S A. 2002. PMID: 12119400 Free PMC article.
-
The coxsackievirus and adenovirus receptor.Curr Top Microbiol Immunol. 2008;323:67-87. doi: 10.1007/978-3-540-75546-3_4. Curr Top Microbiol Immunol. 2008. PMID: 18357766 Review.
-
Picornavirus-receptor interactions.Trends Microbiol. 2002 Jul;10(7):324-31. doi: 10.1016/s0966-842x(02)02383-1. Trends Microbiol. 2002. PMID: 12110211 Review.
Cited by
-
Coxsackievirus B3-Its Potential as an Oncolytic Virus.Viruses. 2021 Apr 21;13(5):718. doi: 10.3390/v13050718. Viruses. 2021. PMID: 33919076 Free PMC article. Review.
-
Receptors and Host Factors for Enterovirus Infection: Implications for Cancer Therapy.Cancers (Basel). 2024 Sep 12;16(18):3139. doi: 10.3390/cancers16183139. Cancers (Basel). 2024. PMID: 39335111 Free PMC article. Review.
-
Structural comparison of different antibodies interacting with parvovirus capsids.J Virol. 2009 Jun;83(11):5556-66. doi: 10.1128/JVI.02532-08. Epub 2009 Mar 25. J Virol. 2009. PMID: 19321620 Free PMC article.
-
A Four-Biomarker Blood Signature Discriminates Systemic Inflammation Due to Viral Infection Versus Other Etiologies.Sci Rep. 2017 Jun 6;7(1):2914. doi: 10.1038/s41598-017-02325-8. Sci Rep. 2017. PMID: 28588308 Free PMC article.
-
Pregnancy and viral infections: Mechanisms of fetal damage, diagnosis and prevention of neonatal adverse outcomes from cytomegalovirus to SARS-CoV-2 and Zika virus.Biochim Biophys Acta Mol Basis Dis. 2021 Oct 1;1867(10):166198. doi: 10.1016/j.bbadis.2021.166198. Epub 2021 Jun 10. Biochim Biophys Acta Mol Basis Dis. 2021. PMID: 34118406 Free PMC article. Review.
References
-
- Bhella, D., I. G. Goodfellow, P. Roversi, Y. Chaudhry, D. J. Evans, and S. M. Lea. 2004. The structure of echovirus type 12 bound to a two-domain fragment of its cellular attachment protein decay-accelerating factor (CD55). J. Biol. Chem. 279:8325-8332. - PubMed
-
- Brodbeck, W. G., D. Liu, J. Sperry, C. Mold, and M. E. Medof. 1996. Localization of classical and alternative pathway regulatory activity within the decay-accelerating factor. J. Immunol. 156:2528-2533. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

