N-myristoylation regulates the SnRK1 pathway in Arabidopsis
- PMID: 17827350
- PMCID: PMC2048702
- DOI: 10.1105/tpc.107.051870
N-myristoylation regulates the SnRK1 pathway in Arabidopsis
Abstract
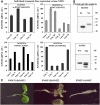
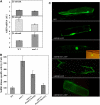
Cotranslational and posttranslational modifications are increasingly recognized as important in the regulation of numerous essential cellular functions. N-myristoylation is a lipid modification ensuring the proper function and intracellular trafficking of proteins involved in many signaling pathways. Arabidopsis thaliana, like human, has two tightly regulated N-myristoyltransferase (NMT) genes, NMT1 and NMT2. Characterization of knockout mutants showed that NMT1 was strictly required for plant viability, whereas NMT2 accelerated flowering. NMT1 impairment induced extremely severe defects in the shoot apical meristem during embryonic development, causing growth arrest after germination. A transgenic plant line with an inducible NMT1 gene demonstrated that NMT1 expression had further effects at later stages. NMT2 did not compensate for NMT1 in the nmt1-1 mutant, but NMT2 overexpression resulted in shoot and root meristem abnormalities. Various data from complementation experiments in the nmt1-1 background, using either yeast or human NMTs, demonstrated a functional link between the developmental arrest of nmt1-1 mutants and the myristoylation state of an extremely small set of protein targets. We show here that protein N-myristoylation is systematically associated with shoot meristem development and that SnRK1 (for SNF1-related kinase) is one of its essential primary targets.
Figures




Similar articles
-
Suppressors of nmtl-181, a conditional lethal allele of the Saccharomyces cerevisiae myristoyl-CoA:protein N-myristoyltransferase gene, reveal proteins involved in regulating protein N-myristoylation.Proc Natl Acad Sci U S A. 1994 Oct 11;91(21):10158-62. doi: 10.1073/pnas.91.21.10158. Proc Natl Acad Sci U S A. 1994. PMID: 7937855 Free PMC article.
-
A role for Saccharomyces cerevisiae fatty acid activation protein 4 in regulating protein N-myristoylation during entry into stationary phase.J Biol Chem. 1998 Oct 2;273(40):25864-74. doi: 10.1074/jbc.273.40.25864. J Biol Chem. 1998. PMID: 9748261
-
Transcription of INO2 and INO4 is regulated by the state of protein N-myristoylation in Saccharomyces cerevisiae.Nucleic Acids Res. 1998 Jun 15;26(12):2865-72. doi: 10.1093/nar/26.12.2865. Nucleic Acids Res. 1998. PMID: 9611229 Free PMC article.
-
[Role of TSK gene in plant shoot and root apical meristem].Tanpakushitsu Kakusan Koso. 2002 Sep;47(12 Suppl):1562-3. Tanpakushitsu Kakusan Koso. 2002. PMID: 12357612 Review. Japanese. No abstract available.
-
N-myristoyltransferase in the leukocytic development processes.Cell Tissue Res. 2011 Aug;345(2):203-11. doi: 10.1007/s00441-011-1202-x. Epub 2011 Jun 24. Cell Tissue Res. 2011. PMID: 21698528 Free PMC article. Review.
Cited by
-
Golgi traffic and integrity depend on N-myristoyl transferase-1 in Arabidopsis.Plant Cell. 2013 May;25(5):1756-73. doi: 10.1105/tpc.113.111393. Epub 2013 May 14. Plant Cell. 2013. PMID: 23673980 Free PMC article.
-
The putative myristoylome of Physcomitrium patens reveals conserved features of myristoylation in basal land plants.Plant Cell Rep. 2023 Jun;42(6):1107-1124. doi: 10.1007/s00299-023-03016-7. Epub 2023 Apr 13. Plant Cell Rep. 2023. PMID: 37052714
-
A novel role for Arabidopsis CBL1 in affecting plant responses to glucose and gibberellin during germination and seedling development.PLoS One. 2013;8(2):e56412. doi: 10.1371/journal.pone.0056412. Epub 2013 Feb 20. PLoS One. 2013. PMID: 23437128 Free PMC article.
-
Cotranslational proteolysis dominates glutathione homeostasis to support proper growth and development.Plant Cell. 2009 Oct;21(10):3296-314. doi: 10.1105/tpc.109.069757. Epub 2009 Oct 23. Plant Cell. 2009. PMID: 19855051 Free PMC article.
-
Beta-subunits of the SnRK1 complexes share a common ancestral function together with expression and function specificities; physical interaction with nitrate reductase specifically occurs via AKINbeta1-subunit.Plant Physiol. 2008 Nov;148(3):1570-82. doi: 10.1104/pp.108.123026. Epub 2008 Sep 3. Plant Physiol. 2008. PMID: 18768910 Free PMC article.
References
-
- Arabidopsis Genome Initiative (2000). Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408 796–815. - PubMed
-
- Ashrafi, K., Farazi, T.A., and Gordon, J.I. (1998). A role for Saccharomyces cerevisiae fatty acid activation protein E in regulating N-myristoylation during entry into stationary phase. J. Biol. Chem. 273 25864–25874. - PubMed
-
- Beeckman, T., and Viane, R. (1999). Embedding thin plant specimens for oriented sectioning. Biotech. Histochem. 75 23–26. - PubMed
-
- Bhatnagar, R.S., Ashrafi, K., Futterer, K., Waksman, G., and Gordon, J.I. (2001). Biology and enzymology of protein N-myristoylation. In The Enzymes, F. Tamanoi and D.S. Sigman, eds (San Diego, CA: Academic Press), pp. 241–286.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

