Role of human HGFIN/nmb in breast cancer
- PMID: 17845721
- PMCID: PMC2242655
- DOI: 10.1186/bcr1764
Role of human HGFIN/nmb in breast cancer
Abstract
Introduction: HGFIN, previously identified as nmb, and its homolog osteoactivin are single transmembrane proteins that are expressed in differentiated immune cells. These proteins exhibit properties that could potentiate tumorigenesis or decrease invasiveness. These seemingly opposing roles of HGFIN suggest that this protein might be central to malignancies and might also behave as a tumor suppressor. Consistent with the reported roles for HGFIN is the fact that this gene is regulated by p53 through multiple binding sites in the 5' flanking region, and is expressed in osteoblasts.
Methods: This study used siRNA to knock-out HGFIN in non-tumorigenic breast cells and ectopically expressed HGFIN in breast cancer cells. In addition, in situ hybridization studies analyzed primary breast tissues from archived breast surgeries. Reporter gene assays studied the untranslated exon 1 of HGFIN.
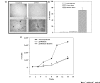
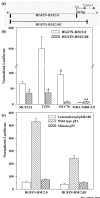
Results: HGFIN expression led to reduced cell growth of breast cancer cells and reduced migration. At the molecular level, reporter gene analyses determined the untranslated exon 1 to be a negative regulator of the upstream enhancing effect. Ectopic expression of wild-type p53 in breast cancer cells that expressed endogenous mutant p53 resulted in increased HGFIN reporter gene activities.
Conclusion: As the majority of cancer cells have mutations in p53, further studies on the relationship between p53 and HGFIN expression, and its role in tumor genesis and bone invasion, might uncover novel therapy targets for breast and other cancers. The results show a central role for p53 in HGFIN expression, which appears to determine the behavior of the cancer cells.
Figures




Comment in
-
Osteoactivin/HGFIN: is it a tumor suppressor or mediator of metastasis in breast cancer?Breast Cancer Res. 2007;9(6):403. doi: 10.1186/bcr1791. Breast Cancer Res. 2007. PMID: 18086324 Free PMC article. No abstract available.
References
-
- Bandari PS, Qian J, Yehia G, Joshi DD, Maloof PB, Potian J, Oh HS, Gascon P, Harrison JS, Rameshwar P. Hematopoietic growth factor inducible neurokinin-1 type: A transmembrane protein that is similar to neurokinin 1 interacts with substance P. Regl Pept. 2003;111:169–178. doi: 10.1016/S0167-0115(02)00288-4. - DOI - PubMed
-
- Ogawa T, Nikawa T, Furochi H, Kosyoji M, Hirasaka K, Suzue N, Sairyo K, Nakano S, Yamaoka T, Itakura M, et al. Osteoactivin upregulates expression of MMP-3 and MMP-9 in fibroblasts infiltrated into denervated skeletal muscle in mice. Am J Physiol Cell Physiol. 2005;289:C697–C707. doi: 10.1152/ajpcell.00565.2004. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

