Human ESCRT and ALIX proteins interact with proteins of the midbody and function in cytokinesis
- PMID: 17853893
- PMCID: PMC2230844
- DOI: 10.1038/sj.emboj.7601850
Human ESCRT and ALIX proteins interact with proteins of the midbody and function in cytokinesis
Erratum in
- EMBO J. 2012 Jul 18;31(14):3228
Abstract
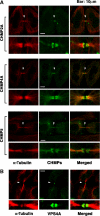
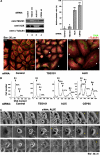
TSG101 and ALIX both function in HIV budding and in vesicle formation at the multivesicular body (MVB), where they interact with other Endosomal Sorting Complex Required for Transport (ESCRT) pathway factors required for release of viruses and vesicles. Proteomic analyses revealed that ALIX and TSG101/ESCRT-I also bind a series of proteins involved in cytokinesis, including CEP55, CD2AP, ROCK1, and IQGAP1. ALIX and TSG101 concentrate at centrosomes and are then recruited to the midbodies of dividing cells through direct interactions between the central CEP55 'hinge' region and GPP-based motifs within TSG101 and ALIX. ESCRT-III and VPS4 proteins are also recruited, indicating that much of the ESCRT pathway localizes to the midbody. Depletion of ALIX and TSG101/ESCRT-I inhibits the abscission step of HeLa cell cytokinesis, as does VPS4 overexpression, confirming a requirement for these proteins in cell division. Furthermore, ALIX point mutants that block CEP55 and CHMP4/ESCRT-III binding also inhibit abscission, indicating that both interactions are essential. These experiments suggest that the ESCRT pathway may be recruited to facilitate analogous membrane fission events during HIV budding, MVB vesicle formation, and the abscission stage of cytokinesis.
Figures






References
-
- Babst M, Katzmann D, Estepa-Sabal E, Meerloo T, Emr S (2002) Escrt-III. An endosome-associated heterooligomeric protein complex required for mvb sorting. Dev Cell 3: 271–282 - PubMed
-
- Bartel PL, Fields BN (1997) The Yeast Two Hybrid Screen. New York: Oxford Press Inc.
-
- Bieniasz PD (2006) Late budding domains and host proteins in enveloped virus release. Virology 344: 55–63 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

