Localization and membrane topology of coronavirus nonstructural protein 4: involvement of the early secretory pathway in replication
- PMID: 17855519
- PMCID: PMC2168994
- DOI: 10.1128/JVI.01506-07
Localization and membrane topology of coronavirus nonstructural protein 4: involvement of the early secretory pathway in replication
Abstract
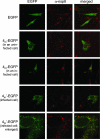
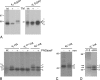
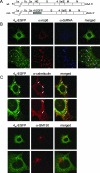
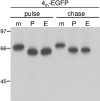
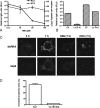
The coronavirus nonstructural proteins (nsp's) derived from the replicase polyproteins collectively constitute the viral replication complexes, which are anchored to double-membrane vesicles. Little is known about the biogenesis of these complexes, the membrane anchoring of which is probably mediated by nsp3, nsp4, and nsp6, as they contain several putative transmembrane domains. As a first step to getting more insight into the formation of the coronavirus replication complex, the membrane topology, processing, and subcellular localization of nsp4 of the mouse hepatitis virus (MHV) and severe acute respiratory syndrome-associated coronavirus (SARS-CoV) were elucidated in this study. Both nsp4 proteins became N glycosylated, while their amino and carboxy termini were localized to the cytoplasm. These observations imply nsp4 to assemble in the membrane as a tetraspanning transmembrane protein with a Nendo/Cendo topology. The amino terminus of SARS-CoV nsp4, but not that of MHV nsp4, was shown to be (partially) processed by signal peptidase. nsp4 localized to the endoplasmic reticulum (ER) when expressed alone but was recruited to the replication complexes in infected cells. nsp4 present in these complexes did not colocalize with markers of the ER or Golgi apparatus, while the susceptibility of its sugars to endoglycosidase H indicated that the protein had also not traveled trough the latter compartment. The important role of the early secretory pathway in formation of the replication complexes was also demonstrated by the inhibition of coronaviral replication when the ER export machinery was blocked by use of the kinase inhibitor H89 or by expression of a mutant, Sar1[H79G].
Figures
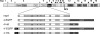


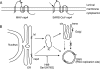
Similar articles
-
Expression and Cleavage of Middle East Respiratory Syndrome Coronavirus nsp3-4 Polyprotein Induce the Formation of Double-Membrane Vesicles That Mimic Those Associated with Coronaviral RNA Replication.mBio. 2017 Nov 21;8(6):e01658-17. doi: 10.1128/mBio.01658-17. mBio. 2017. PMID: 29162711 Free PMC article.
-
Topology and membrane anchoring of the coronavirus replication complex: not all hydrophobic domains of nsp3 and nsp6 are membrane spanning.J Virol. 2008 Dec;82(24):12392-405. doi: 10.1128/JVI.01219-08. Epub 2008 Oct 8. J Virol. 2008. PMID: 18842706 Free PMC article.
-
Mutations across murine hepatitis virus nsp4 alter virus fitness and membrane modifications.J Virol. 2015 Feb;89(4):2080-9. doi: 10.1128/JVI.02776-14. Epub 2014 Dec 3. J Virol. 2015. PMID: 25473044 Free PMC article.
-
The PRRSV replicase: exploring the multifunctionality of an intriguing set of nonstructural proteins.Virus Res. 2010 Dec;154(1-2):61-76. doi: 10.1016/j.virusres.2010.07.030. Epub 2010 Aug 7. Virus Res. 2010. PMID: 20696193 Free PMC article. Review.
-
Insights into RNA synthesis, capping, and proofreading mechanisms of SARS-coronavirus.Virus Res. 2014 Dec 19;194:90-9. doi: 10.1016/j.virusres.2014.10.008. Epub 2014 Oct 17. Virus Res. 2014. PMID: 25451065 Free PMC article. Review.
Cited by
-
Mechanism involved in the pathogenesis and immune response against SARS-CoV-2 infection.Virusdisease. 2021 Jun;32(2):211-219. doi: 10.1007/s13337-021-00687-2. Epub 2021 May 4. Virusdisease. 2021. PMID: 33969150 Free PMC article. Review.
-
Murine hepatitis virus nonstructural protein 4 regulates virus-induced membrane modifications and replication complex function.J Virol. 2010 Jan;84(1):280-90. doi: 10.1128/JVI.01772-09. J Virol. 2010. PMID: 19846526 Free PMC article.
-
Possible Therapeutic Intervention Strategies for COVID-19 by Manipulating the Cellular Proteostasis Network.Adv Exp Med Biol. 2021;1352:125-147. doi: 10.1007/978-3-030-85109-5_8. Adv Exp Med Biol. 2021. PMID: 35132598
-
Murine coronaviruses encoding nsp2 at different genomic loci have altered replication, protein expression, and localization.J Virol. 2008 Dec;82(23):11964-9. doi: 10.1128/JVI.01126-07. Epub 2008 Sep 24. J Virol. 2008. PMID: 18815297 Free PMC article.
-
Biogenesis of cytoplasmic membranous vesicles for plant potyvirus replication occurs at endoplasmic reticulum exit sites in a COPI- and COPII-dependent manner.J Virol. 2008 Dec;82(24):12252-64. doi: 10.1128/JVI.01329-08. Epub 2008 Oct 8. J Virol. 2008. PMID: 18842721 Free PMC article.
References
-
- Altan-Bonnet, N., R. Sougrat, and J. Lippincott-Schwartz. 2004. Molecular basis for Golgi maintenance and biogenesis. Curr. Opin. Cell Biol. 16:364-372. - PubMed
-
- Anand, K., J. Ziebuhr, P. Wadhwani, J. R. Mesters, and R. Hilgenfeld. 2003. Coronavirus main proteinase (3CLpro) structure: basis for design of anti-SARS drugs. Science 300:1763-1767. - PubMed
-
- Aridor, M., and W. E. Balch. 2000. Kinase signaling initiates coat complex II (COPII) recruitment and export from the mammalian endoplasmic reticulum. J. Biol. Chem. 275:35673-35676. - PubMed
-
- Belov, G. A., and E. Ehrenfeld. 2007. Involvement of cellular membrane traffic proteins in poliovirus replication. Cell Cycle 6:36-38. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

