HAP1 can sequester a subset of TBP in cytoplasmic inclusions via specific interaction with the conserved TBP(CORE)
- PMID: 17868456
- PMCID: PMC2082042
- DOI: 10.1186/1471-2199-8-76
HAP1 can sequester a subset of TBP in cytoplasmic inclusions via specific interaction with the conserved TBP(CORE)
Abstract
Background: Huntington's disease, spinal and bulbar muscular atrophy, and spinocerebellar ataxia 17 (SCA17) are caused by expansions in the polyglutamine (polyQ) repeats in Huntingtin protein (Htt), androgen receptor protein (AR), and TATA-binding protein (TBP), respectively. Htt-associated protein 1 (HAP1), a component of neuronal cytoplasmic stigmoid bodies (STBs), can sequester polyQ-expanded Htt and AR in STBs, thereby antagonizing formation of the nuclear aggregates associated with apoptotic neuron loss and disease progression.
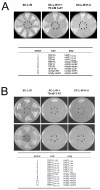
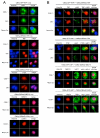
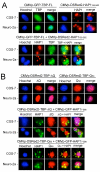
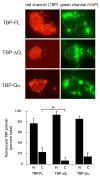
Results: Clones of HAP1 were isolated from unbiased two-hybrid screens for proteins that interact with TBP. Domain mapping showed that regions between amino acids 157 and 261 and between amino acids 473 and 582 of mouse HAP1 both bind specifically to the conserved C-terminal TBP(CORE) domain, away from the TBP N-terminal polyQ region. When fluorescently tagged versions of HAP1 or TBP were expressed independently in COS-7, 293, or Neuro-2a cells, all TBP localized to the nucleus and all HAP1 assembled into cytoplasmic stigmoid-like bodies (STLBs). When co-expressed, a portion of the TBP was assembled into the HAP1 STLBs while the remainder was localized to the nucleus. Although the TBP N terminus, including the polyQ region, was unnecessary for TBP-HAP1 interaction, in mammalian cells, removal of the TBP Q(repeat) reduced the proportion of TBP that assembled into STLBs, whereas expansion of the Q(repeat) had no significant affect on TBP subcellular localization.
Conclusion: HAP1 can sequester a subset of TBP protein away from the nucleus; extranuclear TBP sequestration is quantitatively influenced by the TBP polyQ repeat. These results suggest HAP1 could provide protection from SCA17 neuropathology.
Figures




Similar articles
-
Polyglutamine expansion reduces the association of TATA-binding protein with DNA and induces DNA binding-independent neurotoxicity.J Biol Chem. 2008 Mar 28;283(13):8283-90. doi: 10.1074/jbc.M709674200. Epub 2008 Jan 24. J Biol Chem. 2008. PMID: 18218637 Free PMC article.
-
Molecular investigation of TBP allele length: a SCA17 cellular model and population study.Neurobiol Dis. 2003 Jun;13(1):37-45. doi: 10.1016/s0969-9961(03)00014-7. Neurobiol Dis. 2003. PMID: 12758065
-
Deactivation of TBP contributes to SCA17 pathogenesis.Hum Mol Genet. 2014 Dec 20;23(25):6878-93. doi: 10.1093/hmg/ddu410. Epub 2014 Aug 7. Hum Mol Genet. 2014. PMID: 25104854
-
Proteomics of polyglutamine aggregates.Methods Enzymol. 2006;412:63-76. doi: 10.1016/S0076-6879(06)12005-4. Methods Enzymol. 2006. PMID: 17046652 Review.
-
The selective vulnerability of nerve cells in Huntington's disease.Neuropathol Appl Neurobiol. 2001 Feb;27(1):1-21. doi: 10.1046/j.0305-1846.2001.00299.x. Neuropathol Appl Neurobiol. 2001. PMID: 11298997 Review.
Cited by
-
Huntingtin associated protein 1 and its functions.Cell Adh Migr. 2009 Jan-Mar;3(1):71-6. doi: 10.4161/cam.3.1.7511. Epub 2009 Jan 26. Cell Adh Migr. 2009. PMID: 19262167 Free PMC article. Review.
-
Huntingtin mediates dendritic transport of β-actin mRNA in rat neurons.Sci Rep. 2011;1:140. doi: 10.1038/srep00140. Epub 2011 Nov 3. Sci Rep. 2011. PMID: 22355657 Free PMC article.
-
Microtubule-dependent formation of the stigmoid body as a cytoplasmic inclusion distinct from pathological aggresomes.Histochem Cell Biol. 2009 Sep;132(3):305-18. doi: 10.1007/s00418-009-0618-9. Epub 2009 Jul 4. Histochem Cell Biol. 2009. PMID: 19578869
-
The adaptor proteins HAP1a and GRIP1 collaborate to activate the kinesin-1 isoform KIF5C.J Cell Sci. 2019 Dec 13;132(24):jcs215822. doi: 10.1242/jcs.215822. J Cell Sci. 2019. PMID: 31757889 Free PMC article.
-
Biological functions and potential therapeutic applications of huntingtin-associated protein 1: progress and prospects.Clin Transl Oncol. 2022 Feb;24(2):203-214. doi: 10.1007/s12094-021-02702-w. Epub 2021 Sep 26. Clin Transl Oncol. 2022. PMID: 34564830 Review.
References
-
- MacDonald ME, Ambrose CM, Duyao MP, Myers RH, Lin C, Srinidhi L, Barnes G, Taylor SA, James M, Groot N, MacFarlane H, Jenkins B, Anderson MA, Wexler NS, Gusella JF, Bates GP, Baxendale S, Hummerich H, Kirby S, North M, Youngman S, Mott R, Zehetner G, Sedlacek Z, Poustka A, Frischauf A-M, Lehrach H, Buckler AJ, Church D, Doucette-Stamm L, et al. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. The Huntington's Disease Collaborative Research Group. Cell. 1993;72:971–983. doi: 10.1016/0092-8674(93)90585-E. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

