Biochemical characterization of a SET and transposase fusion protein, Metnase: its DNA binding and DNA cleavage activity
- PMID: 17877369
- PMCID: PMC3374406
- DOI: 10.1021/bi7005477
Biochemical characterization of a SET and transposase fusion protein, Metnase: its DNA binding and DNA cleavage activity
Abstract
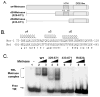
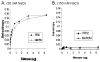
Metnase (SETMAR) is a SET and transposase fusion protein that promotes in vivo end joining activity and mediates genomic integration of foreign DNA. Recent studies showed that Metnase retained most of the transposase activities, including 5'-terminal inverted repeat (TIR)-specific binding and assembly of a paired end complex, and cleavage of the 5'-end of the TIR element. Here we show that R432 within the helix-turn-helix motif is critical for sequence-specific recognition, as the R432A mutation abolishes its TIR-specific DNA binding activity. Metnase possesses a unique DNA nicking and/or endonuclease activity that mediates cleavage of duplex DNA in the absence of the TIR sequence. While the HTH motif is essential for the Metnase-TIR interaction, it is not required for its DNA cleavage activity. The DDE-like motif is crucial for its DNA cleavage action as a point mutation at this motif (D483A) abolished its DNA cleavage activity. Together, our results suggest that Metnase's DNA cleavage activity, unlike those of other eukaryotic transposases, is not coupled to its sequence-specific DNA binding.
Figures






References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

