Back to basics: the untreated rabbit reticulocyte lysate as a competitive system to recapitulate cap/poly(A) synergy and the selective advantage of IRES-driven translation
- PMID: 17881372
- PMCID: PMC2094066
- DOI: 10.1093/nar/gkm682
Back to basics: the untreated rabbit reticulocyte lysate as a competitive system to recapitulate cap/poly(A) synergy and the selective advantage of IRES-driven translation
Abstract
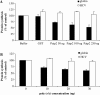
Translation of most eukaryotic mRNAs involves the synergistic action between the 5' cap structure and the 3' poly(A) tail at the initiation step. The poly(A) tail has also been shown to stimulate translation of picornavirus internal ribosome entry sites (IRES)-directed translation. These effects have been attributed principally to interactions between eIF4G and poly(A)-binding protein (PABP) but also to the participation of PABP in other steps during translation initiation. As the rabbit reticulocyte lysate (RRL) does not recapitulate this cap/poly(A) synergy, several systems based on cellular cell-free extracts have been developed to study the effects of poly(A) tail in vitro but they generally exhibit low translational efficiency. Here, we describe that the non-nuclease-treated RRL (untreated RRL) is able to recapitulate the effects of poly(A) tail on translation in vitro. In this system, translation of a capped/polyadenylated RNA was specifically inhibited by either Paip2 or poly(rA), whereas translation directed by HCV IRES remained unaffected. Moreover, cleavage of eIF4G by FMDV L protease strongly stimulated translation directed by the EMCV IRES, thus recapitulating the competitive advantage that the proteolytic processing of eIF4G confers to IRES-driven RNAs.
Figures



Similar articles
-
Cap-Poly(A) synergy in mammalian cell-free extracts. Investigation of the requirements for poly(A)-mediated stimulation of translation initiation.J Biol Chem. 2000 Oct 13;275(41):32268-76. doi: 10.1074/jbc.M004304200. J Biol Chem. 2000. PMID: 10922367
-
Eukaryotic initiation factor 4G-poly(A) binding protein interaction is required for poly(A) tail-mediated stimulation of picornavirus internal ribosome entry segment-driven translation but not for X-mediated stimulation of hepatitis C virus translation.Mol Cell Biol. 2001 Jul;21(13):4097-109. doi: 10.1128/MCB.21.13.4097-4109.2001. Mol Cell Biol. 2001. PMID: 11390639 Free PMC article.
-
Biochemical characterisation of cap-poly(A) synergy in rabbit reticulocyte lysates: the eIF4G-PABP interaction increases the functional affinity of eIF4E for the capped mRNA 5'-end.Nucleic Acids Res. 2000 Nov 1;28(21):4068-75. doi: 10.1093/nar/28.21.4068. Nucleic Acids Res. 2000. PMID: 11058101 Free PMC article.
-
[Translational control by the poly(A) binding protein: a check for mRNA integrity].Mol Biol (Mosk). 2006 Jul-Aug;40(4):684-93. Mol Biol (Mosk). 2006. PMID: 16913227 Review. Russian.
-
RNA-protein interactions in regulation of picornavirus RNA translation.Microbiol Rev. 1996 Sep;60(3):499-511. doi: 10.1128/mr.60.3.499-511.1996. Microbiol Rev. 1996. PMID: 8840784 Free PMC article. Review.
Cited by
-
A researcher's guide to the galaxy of IRESs.Cell Mol Life Sci. 2017 Apr;74(8):1431-1455. doi: 10.1007/s00018-016-2409-5. Epub 2016 Nov 16. Cell Mol Life Sci. 2017. PMID: 27853833 Free PMC article. Review.
-
Epstein-Barr Virus Protein EB2 Stimulates Translation Initiation of mRNAs through Direct Interactions with both Poly(A)-Binding Protein and Eukaryotic Initiation Factor 4G.J Virol. 2018 Jan 17;92(3):e01917-17. doi: 10.1128/JVI.01917-17. Print 2018 Feb 1. J Virol. 2018. PMID: 29142127 Free PMC article.
-
A structural element within the 5'UTR of β-catenin mRNA modulates its translation under hypoxia.Nucleic Acids Res. 2025 Apr 22;53(8):gkaf321. doi: 10.1093/nar/gkaf321. Nucleic Acids Res. 2025. PMID: 40309781 Free PMC article.
-
mRNA structural elements immediately upstream of the start codon dictate dependence upon eIF4A helicase activity.Genome Biol. 2019 Dec 30;20(1):300. doi: 10.1186/s13059-019-1901-2. Genome Biol. 2019. PMID: 31888698 Free PMC article.
-
DEAD-box protein DDX3 associates with eIF4F to promote translation of selected mRNAs.EMBO J. 2012 Sep 12;31(18):3745-56. doi: 10.1038/emboj.2012.220. Epub 2012 Aug 7. EMBO J. 2012. PMID: 22872150 Free PMC article.
References
-
- Gingras AC, Raught B, Sonenberg N. eIF4 initiation factors: effectors of mRNA recruitment to ribosomes and regulators of translation. Annu. Rev. Biochem. 1999;68:913–963. - PubMed
-
- Gallie DR. The cap and poly(A) tail function synergistically to regulate mRNA translational efficiency. Genes. Dev. 1991;5:2108–2116. - PubMed
-
- Prevot D, Darlix JL, Ohlmann T. Conducting the initiation of protein synthesis: the role of eIF4G. Biol. Cell. 2003;95:141–156. - PubMed
-
- Pelletier J, Sonenberg N. Internal initiation of translation of eukaryotic mRNA directed by a sequence derived from poliovirus RNA. Nature. 1988;334:320–325. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

