Endocytosis of DNA-Hsp65 alters the pH of the late endosome/lysosome and interferes with antigen presentation
- PMID: 17895965
- PMCID: PMC1976595
- DOI: 10.1371/journal.pone.0000923
Endocytosis of DNA-Hsp65 alters the pH of the late endosome/lysosome and interferes with antigen presentation
Abstract
Background: Experimental models using DNA vaccine has shown that this vaccine is efficient in generating humoral and cellular immune responses to a wide variety of DNA-derived antigens. Despite the progress in DNA vaccine development, the intracellular transport and fate of naked plasmid DNA in eukaryotic cells is poorly understood, and need to be clarified in order to facilitate the development of novel vectors and vaccine strategies.
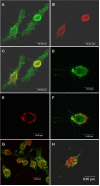
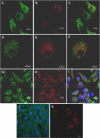
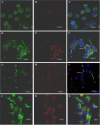
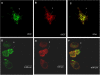
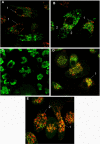
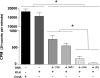
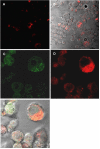
Methodology and principal findings: Using confocal microscopy, we have demonstrated for the first time that after plasmid DNA uptake an inhibition of the acidification of the lysosomal compartment occurs. This lack of acidification impaired antigen presentation to CD4 T cells, but did not alter the recruitment of MyD88. The recruitment of Rab 5 and Lamp I were also altered since we were not able to co-localize plasmid DNA with Rab 5 and Lamp I in early endosomes and late endosomes/lysosomes, respectively. Furthermore, we observed that the DNA capture process in macrophages was by clathrin-mediated endocytosis. In addition, we observed that plasmid DNA remains in vesicles until it is in a juxtanuclear location, suggesting that the plasmid does not escape into the cytoplasmic compartment.
Conclusions and significance: Taken together our data suggests a novel mechanism involved in the intracellular trafficking of plasmid DNA, and opens new possibilities for the use of lower doses of plasmid DNA to regulate the immune response.
Conflict of interest statement
Figures
References
-
- Gurunathan S, Klinman DM, Seder, RA DNA vaccines: immunology, application, and optimization. Annu Rev Immunol. 2000;18:927–974. - PubMed
-
- Wolff JA, Malone RW, Williams P, Chong W, Acsadi G, et al. Direct gene transfer into mouse muscle in vivo. Science. 1990;247:1465–1468. - PubMed
-
- Acsadi G, Jiao SS, Jani A, Duke D, Williams P, et al. Direct gene transfer and expression into rat heart in vivo. New Biol. 1991;3:71–81. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

