Mammalian microRNA prediction through a support vector machine model of sequence and structure
- PMID: 17895987
- PMCID: PMC1978525
- DOI: 10.1371/journal.pone.0000946
Mammalian microRNA prediction through a support vector machine model of sequence and structure
Abstract
Background: MicroRNAs (miRNAs) are endogenous small noncoding RNA gene products, on average 22 nt long, found in a wide variety of organisms. They play important regulatory roles by targeting mRNAs for degradation or translational repression. There are 377 known mouse miRNAs and 475 known human miRNAs in the May 2007 release of the miRBase database, the majority of which are conserved between the two species. A number of recent reports imply that it is likely that many mammalian miRNAs remain to be discovered. The possibility that there are more of them expressed at lower levels or in more specialized expression contexts calls for the exploitation of genome sequence information to accelerate their discovery.
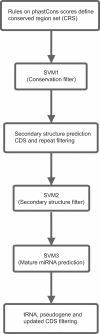
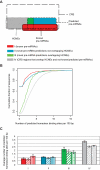
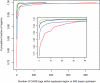
Methodology/principal findings: In this article, we describe a computational method-mirCoS-that uses three support vector machine models sequentially to discover new miRNA candidates in mammalian genomes based on sequence, secondary structure, and conservation. mirCoS can efficiently detect the majority of known miRNAs and predicts an extensive set of hairpin structures based on human-mouse comparisons. In total, 3476 mouse candidates and 3441 human candidates were found. These hairpins are more similar to known miRNAs than to negative controls in several aspects not considered by the prediction algorithm. A significant fraction of predictions is supported by existing expression evidence.
Conclusions/significance: Using a novel approach, mirCoS performs comparably to or better than existing miRNA prediction methods, and contributes a significant number of new candidate miRNAs for experimental verification.
Conflict of interest statement
Figures


Similar articles
-
Classification of real and pseudo microRNA precursors using local structure-sequence features and support vector machine.BMC Bioinformatics. 2005 Dec 29;6:310. doi: 10.1186/1471-2105-6-310. BMC Bioinformatics. 2005. PMID: 16381612 Free PMC article.
-
Bioinformatic discovery of microRNA precursors from human ESTs and introns.BMC Genomics. 2006 Jul 3;7:164. doi: 10.1186/1471-2164-7-164. BMC Genomics. 2006. PMID: 16813663 Free PMC article.
-
Analysis of microRNA-target interactions by a target structure based hybridization model.Pac Symp Biocomput. 2008:64-74. Pac Symp Biocomput. 2008. PMID: 18232104
-
Prediction of human microRNA targets.Methods Mol Biol. 2006;342:101-13. doi: 10.1385/1-59745-123-1:101. Methods Mol Biol. 2006. PMID: 16957370 Review.
-
Computational identification of microRNAs and their targets.Birth Defects Res C Embryo Today. 2006 Jun;78(2):118-28. doi: 10.1002/bdrc.20067. Birth Defects Res C Embryo Today. 2006. PMID: 16847881 Review.
Cited by
-
MicroRNA: Implications for Alzheimer Disease and other Human CNS Disorders.Curr Genomics. 2009 May;10(3):154-68. doi: 10.2174/138920209788185252. Curr Genomics. 2009. PMID: 19881909 Free PMC article.
-
The effect of central loops in miRNA:MRE duplexes on the efficiency of miRNA-mediated gene regulation.PLoS One. 2008 Mar 5;3(3):e1719. doi: 10.1371/journal.pone.0001719. PLoS One. 2008. PMID: 18320040 Free PMC article.
-
Bioinformatic tools for microRNA dissection.Nucleic Acids Res. 2016 Jan 8;44(1):24-44. doi: 10.1093/nar/gkv1221. Epub 2015 Nov 17. Nucleic Acids Res. 2016. PMID: 26578605 Free PMC article. Review.
-
Small RNA expression profiling by high-throughput sequencing: implications of enzymatic manipulation.J Nucleic Acids. 2012;2012:360358. doi: 10.1155/2012/360358. Epub 2012 Jun 20. J Nucleic Acids. 2012. PMID: 22778911 Free PMC article.
-
Small RNAs in Plant Responses to Abiotic Stresses: Regulatory Roles and Study Methods.Int J Mol Sci. 2015 Oct 15;16(10):24532-54. doi: 10.3390/ijms161024532. Int J Mol Sci. 2015. PMID: 26501263 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

