The PDZ domain as a complex adaptive system
- PMID: 17895993
- PMCID: PMC1978516
- DOI: 10.1371/journal.pone.0000953
The PDZ domain as a complex adaptive system
Abstract
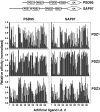
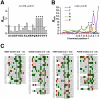
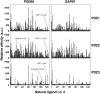
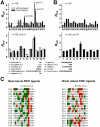
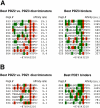
Specific protein associations define the wiring of protein interaction networks and thus control the organization and functioning of the cell as a whole. Peptide recognition by PDZ and other protein interaction domains represents one of the best-studied classes of specific protein associations. However, a mechanistic understanding of the relationship between selectivity and promiscuity commonly observed in the interactions mediated by peptide recognition modules as well as its functional meaning remain elusive. To address these questions in a comprehensive manner, two large populations of artificial and natural peptide ligands of six archetypal PDZ domains from the synaptic proteins PSD95 and SAP97 were generated by target-assisted iterative screening (TAIS) of combinatorial peptide libraries and by synthesis of proteomic fragments, correspondingly. A comparative statistical analysis of affinity-ranked artificial and natural ligands yielded a comprehensive picture of known and novel PDZ ligand specificity determinants, revealing a hitherto unappreciated combination of specificity and adaptive plasticity inherent to PDZ domain recognition. We propose a reconceptualization of the PDZ domain in terms of a complex adaptive system representing a flexible compromise between the rigid order of exquisite specificity and the chaos of unselective promiscuity, which has evolved to mediate two mutually contradictory properties required of such higher order sub-cellular organizations as synapses, cell junctions, and others--organizational structure and organizational plasticity/adaptability. The generalization of this reconceptualization in regard to other protein interaction modules and specific protein associations is consistent with the image of the cell as a complex adaptive macromolecular system as opposed to clockwork.
Conflict of interest statement
Figures
References
-
- Kuriyan J, Cowburn D. Modular peptide recognition domains in eukaryotic signaling. Annu Rev Biophys Biomol Struct. 1997;26:259–288. - PubMed
-
- Sudol M. From Src Homology domains to other signaling modules: proposal of the ‘protein recognition code’. Oncogene. 1998;17:1469–1474. - PubMed
-
- Castagnoli L, Costantini A, Dall'Armi C, Gonfloni S, Montecchi-Palazzi L, et al. Selectivity and promiscuity in the interaction network mediated by protein recognition modules. FEBS Lett. 2004;567:74–79. - PubMed
-
- Kay BK, Kasanov J, Knight S, Kurakin A. Convergent evolution with combinatorial peptides. FEBS Lett. 2000;480:55–62. - PubMed
-
- Zucconi A, Panni S, Paoluzi S, Castagnoli L, Dente L, et al. Domain repertoires as a tool to derive protein recognition rules. FEBS Lett. 2000;480:49–54. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

