Virp1 is a host protein with a major role in Potato spindle tuber viroid infection in Nicotiana plants
- PMID: 17898061
- PMCID: PMC2169090
- DOI: 10.1128/JVI.00974-07
Virp1 is a host protein with a major role in Potato spindle tuber viroid infection in Nicotiana plants
Abstract
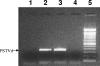
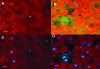
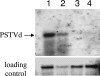
Viroids are small, circular, single-stranded RNA molecules that, while not coding for any protein, cause several plant diseases. Viroids rely for their infectious cycle on host proteins, most of which are likely to be involved in endogenous RNA-mediated phenomena. Therefore, characterization of host factors interacting with the viroid may contribute to the elucidation of RNA-related pathways of the hosts. Potato spindle tuber viroid (PSTVd) infects several members of the Solanaceae family. In an RNA ligand screening we have previously isolated the tomato protein Virp1 by its ability to specifically interact with PSTVd positive-strand RNA. Virp1 is a bromodomain-containing protein with an atypical RNA binding domain and a nuclear localization signal. Here we investigate the role of Virp1 in the viroid infection cycle by the use of transgenic lines of Nicotiana tabacum and Nicotiana benthamiana that either overexpress the tomato Virp1 RNA or suppress the orthologous Nicotiana genes through RNA silencing. Plants of the Virp1-suppressed lines were not infected by PSTVd or Citrus exocortis viroid through mechanical inoculation, indicating a major role of Virp1 in viroid infection. On the other hand, overexpression of tomato Virp1 in N. tabacum and N. benthamiana plants did not affect PSTVd KF 440-2 infectivity or symptomatology in these species. Transfection experiments with isolated protoplasts revealed that Virp1-suppressed cells were unable to sustain viroid replication, suggesting that resistance to viroid infection in Virp1-suppressed plants is likely the result of cell-autonomous events.
Figures





References
-
- Branch, A. D., and H. D. Robertson. 1984. A replication cycle for viroids and other small infectious RNA's. Science 223:450-455. - PubMed
-
- Denti, M. A., A. Boutla, M. Tsagris, and M. Tabler. 2004. Short interfering RNAs specific for potato spindle tuber viroid are found in the cytoplasm but not in the nucleus. Plant J. 37:762-769. - PubMed
-
- Dhalluin, C., J. E. Carlson, L. Zeng, C. He, A. K. Aggarwal, and M. M. Zhou. 1999. Structure and ligand of a histone acetyltransferase bromodomain. Nature 399:491-496. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

