The fluxes through glycolytic enzymes in Saccharomyces cerevisiae are predominantly regulated at posttranscriptional levels
- PMID: 17898166
- PMCID: PMC2000426
- DOI: 10.1073/pnas.0707476104
The fluxes through glycolytic enzymes in Saccharomyces cerevisiae are predominantly regulated at posttranscriptional levels
Abstract
Metabolic fluxes may be regulated "hierarchically," e.g., by changes of gene expression that adjust enzyme capacities (V(max)) and/or "metabolically" by interactions of enzymes with substrates, products, or allosteric effectors. In the present study, a method is developed to dissect the hierarchical regulation into contributions by transcription, translation, protein degradation, and posttranslational modification. The method was applied to the regulation of fluxes through individual glycolytic enzymes when the yeast Saccharomyces cerevisiae was confronted with the absence of oxygen and the presence of benzoic acid depleting its ATP. Metabolic regulation largely contributed to the approximately 10-fold change in flux through the glycolytic enzymes. This contribution varied from 50 to 80%, depending on the glycolytic step and the cultivation condition tested. Within the 50-20% hierarchical regulation of fluxes, transcription played a minor role, whereas regulation of protein synthesis or degradation was the most important. These also contributed to 75-100% of the regulation of protein levels.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
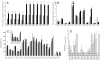
References
-
- Suter B, Auerbach D, Stagljar I. Biotechniques. 2006;40:625–644. - PubMed
-
- Griffin TJ, Gygi SP, Ideker T, Rist B, Eng J, Hood L, Aebersold R. Mol Cell Proteomics. 2002;1:323–333. - PubMed
-
- Daran-Lapujade P, Jansen MLA, Daran JM, van Gulik W, de Winde JH, Pronk JT. J Biol Chem. 2004;278:3265–3274.
-
- Yang C, Hua Q, Shimizu K. Appl Microbiol Biotechnol. 2002;58:813–822. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

