Limited genetic diversity in Salmonella enterica serovar Enteritidis PT13
- PMID: 17908316
- PMCID: PMC2137926
- DOI: 10.1186/1471-2180-7-87
Limited genetic diversity in Salmonella enterica serovar Enteritidis PT13
Abstract
Background: Salmonella enterica serovar Enteritidis has emerged as a significant foodborne pathogen throughout the world and is commonly characterized by phage typing. In Canada phage types (PT) 4, 8 and 13 predominate and in 2005 a large foodborne PT13 outbreak occurred in the province of Ontario. The ability to link strains during this outbreak was difficult due to the apparent clonality of PT13 isolates in Canada, as there was a single dominant pulsed-field gel electrophoresis (PFGE) profile amongst epidemiologically linked human and food isolates as well as concurrent sporadic strains. The aim of this study was to perform comparative genomic hybridization (CGH), DNA sequence-based typing (SBT) genomic analyses, plasmid analyses, and automated repetitive sequence-based PCR (rep-PCR) to identify epidemiologically significant traits capable of subtyping S. Enteritidis PT13.
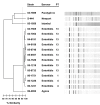
Results: CGH using an oligonucleotide array based upon chromosomal coding sequences of S. enterica serovar Typhimurium strain LT2 and the Salmonella genomic island 1 successfully determined major genetic differences between S. Typhimurium and S. Enteritidis PT13, but no significant strain-to-strain differences were observed between S. Enteritidis PT13 isolates. Individual loci (safA and fliC) that were identified as potentially divergent in the CGH data set were sequenced in a panel of S. Enteritidis strains, and no differences were detected between the PT13 strains. Additional sequence-based typing was performed at the fimA, mdh, manB, cyaA, citT, caiC, dmsA, ratA and STM0660 loci. Similarly, no diversity was observed amongst PT13 strains. Variation in plasmid content between PT13 strains was observed, but macrorestriction with BglII did not identify further differences. Automated rep-PCR patterns were variable between serovars, but S. Enteritidis PT13 strains could not be differentiated.
Conclusion: None of the methods identified any significant variation between PT13 strains. Greater than 11,300 base pairs of sequence for each of seven S. Enteritidis PT13 strains were analyzed without detecting a single polymorphic site, although diversity between different phage types of S. Enteritidis was observed. These data suggest that Canadian S. Enteritidis PT13 strains are highly related genetically.
Figures



References
-
- Poppe C. Epidemiology of Salmonella enterica Serovar Enteritidis. In: Saeed AM, Gast RK, Potter ME and Wall PG, editor. Salmonella enterica Serovar Enteritidis in Humans and Animals. Ames, Iowa State University Press; 1999. pp. 3–18.
-
- Munro DS, Girdwood RWA, Reilly WJ. Salmonella enterica Serovar Enteritidis in Scotland. In: Saeed AM, Gast RK, Potter ME and Wall PG, editor. Salmonella enterica Serovar Enteritidis in Humans and Animals. Ames, Iowa State University Press; 1999. pp. 27–31.
-
- Tschape H, Liesegang A, Gericke B, Prager R, Rabsch W, Helmuth R. Ups and Downs of Salmonella enterica Serovar Enteritidis in Germany. In: Saeed AM, Gast RK, Potter ME and Wall PG, editor. Salmonella enterica Serovar Enteritidis in Humans and Animals. Ames, Iowa State University Press; 1999. pp. 51–61.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

