Functional effects of single nucleotide polymorphisms in the coding region of human N-acetyltransferase 1
- PMID: 17909564
- PMCID: PMC2575040
- DOI: 10.1038/sj.tpj.6500483
Functional effects of single nucleotide polymorphisms in the coding region of human N-acetyltransferase 1
Abstract
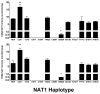
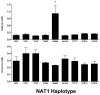
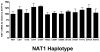
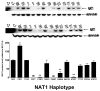
Genetic variants of human N-acetyltransferase 1 (NAT1) are associated with cancer and birth defects. N- and O-acetyltransferase catalytic activities, Michaelis-Menten kinetic constants (K(m) and V(max)) and steady-state expression levels of NAT1-specific mRNA and protein were determined for the reference NAT1*4 and variant human NAT1 haplotypes possessing single nucleotide polymorphisms (SNPs) in the open reading frame. Although none of the SNPs caused a significant effect on steady-state levels of NAT1-specific mRNA, C97T(R33stop), C190T(R64W), C559T (R187stop) and A752T(D251V) each reduced NAT1 protein level and/or N- and O-acetyltransferase catalytic activities to levels below detection. G560A(R187Q) substantially reduced NAT1 protein level and catalytic activities and increased substrate K(m). The G445A(V149I), G459A(synonymous) and T640G(S214A) haplotype present in NAT1*11 significantly (P<0.05) increased NAT1 protein level and catalytic activity. Neither T21G(synonymous), T402C(synonymous), A613G(M205V), T777C(synonymous), G781A(E261K) nor A787G(I263V) significantly affected K(m), catalytic activity, mRNA or protein level. These results suggest heterogeneity among slow NAT1 acetylator phenotypes.
Figures
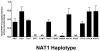
Similar articles
-
Functional effects of genetic polymorphisms in the N-acetyltransferase 1 coding and 3' untranslated regions.Birth Defects Res A Clin Mol Teratol. 2011 Feb;91(2):77-84. doi: 10.1002/bdra.20763. Epub 2011 Feb 2. Birth Defects Res A Clin Mol Teratol. 2011. PMID: 21290563 Free PMC article.
-
Structure-function analyses of single nucleotide polymorphisms in human N-acetyltransferase 1.Drug Metab Rev. 2008;40(1):169-84. doi: 10.1080/03602530701852917. Drug Metab Rev. 2008. PMID: 18259988 Free PMC article. Review.
-
560G>A (rs4986782) (R187Q) Single Nucleotide Polymorphism in Arylamine N-Acetyltransferase 1 Increases Affinity for the Aromatic Amine Carcinogens 4-Aminobiphenyl and N-Hydroxy-4-Aminobiphenyl: Implications for Cancer Risk Assessment.Front Pharmacol. 2022 Feb 22;13:820082. doi: 10.3389/fphar.2022.820082. eCollection 2022. Front Pharmacol. 2022. PMID: 35273499 Free PMC article.
-
Variants of N-acetyltransferase NAT1 and a case-control study of colorectal adenomas.Pharmacogenetics. 1998 Jun;8(3):269-81. doi: 10.1097/00008571-199806000-00009. Pharmacogenetics. 1998. PMID: 9682272
-
Arylamine N-acetyltransferases: structural and functional implications of polymorphisms.Toxicology. 2008 Dec 30;254(3):170-83. doi: 10.1016/j.tox.2008.08.022. Epub 2008 Sep 12. Toxicology. 2008. PMID: 18852012 Review.
Cited by
-
Case Study 10: A Case to Investigate Acetyl Transferase Kinetics.Methods Mol Biol. 2021;2342:781-808. doi: 10.1007/978-1-0716-1554-6_29. Methods Mol Biol. 2021. PMID: 34272717
-
Functional effects of genetic polymorphisms in the N-acetyltransferase 1 coding and 3' untranslated regions.Birth Defects Res A Clin Mol Teratol. 2011 Feb;91(2):77-84. doi: 10.1002/bdra.20763. Epub 2011 Feb 2. Birth Defects Res A Clin Mol Teratol. 2011. PMID: 21290563 Free PMC article.
-
Humans and Chimpanzees Display Opposite Patterns of Diversity in Arylamine N-Acetyltransferase Genes.G3 (Bethesda). 2019 Jul 9;9(7):2199-2224. doi: 10.1534/g3.119.400223. G3 (Bethesda). 2019. PMID: 31068377 Free PMC article.
-
Comparative analysis of xenobiotic metabolising N-acetyltransferases from ten non-human primates as in vitro models of human homologues.Sci Rep. 2018 Jun 27;8(1):9759. doi: 10.1038/s41598-018-28094-6. Sci Rep. 2018. PMID: 29950659 Free PMC article.
-
Functional expression of human arylamine N-acetyltransferase NAT1*10 and NAT1*11 alleles: a mini review.Pharmacogenet Genomics. 2018 Oct;28(10):238-244. doi: 10.1097/FPC.0000000000000350. Pharmacogenet Genomics. 2018. PMID: 30222709 Free PMC article. Review.
References
-
- Butcher NJ, Boukouvala S, Sim E, Minchin RF. Pharmacogenetics of the arylamine N-acetyltransferases. Pharmacogenomics J. 2002;2:30–42. - PubMed
-
- Boukouvala S, Fakis G. Arylamine N-acetyltransferases: what we learn from genes and genomes. Drug Metab Rev. 2005;37:511–64. - PubMed
-
- Hein DW. Molecular genetics and function of NAT1 and NAT2: role in aromatic amine metabolism and carcinogenesis. Mutat Res. 2002;506–507:65–77. - PubMed
-
- Hein DW, Doll MA, Fretland AJ, Leff MA, Webb SJ, Xiao GH, et al. Molecular genetics and epidemiology of the NAT1 and NAT2 acetylation polymorphisms. Cancer Epidemiol Biomarkers Prev. 2000;9:29–42. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

