Natural killer T-cell characterization through gene expression profiling: an account of versatility bridging T helper type 1 (Th1), Th2 and Th17 immune responses
- PMID: 17916165
- PMCID: PMC2433278
- DOI: 10.1111/j.1365-2567.2007.02701.x
Natural killer T-cell characterization through gene expression profiling: an account of versatility bridging T helper type 1 (Th1), Th2 and Th17 immune responses
Abstract
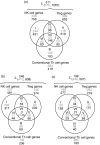
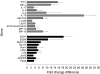
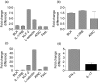
Natural killer T (NKT) cells constitute a distinct lymphocyte lineage at the interface between innate and adaptive immunity, yet their role in the immune response remains elusive. Whilst NKT cells share features with other conventional T lymphocytes, they are unique in their rapid, concomitant production of T helper type 1 (Th1) and Th2 cytokines upon T-cell receptor (TCR) ligation. In order to characterize the gene expression of NKT cells, we performed comparative microarray analyses of murine resting NKT cells, natural killer (NK) cells and naïve conventional CD4+ T helper (Th) and regulatory T cells (Treg). We then compared the gene expression profiles of resting and alpha-galactosylceramide (alphaGalCer)-activated NKT cells to elucidate the gene expression signature upon activation. We describe here profound differences in gene expression among the various cell types and the identification of a unique NKT cell gene expression profile. In addition to known NKT cell-specific markers, many genes were expressed in NKT cells that had not been attributed to this population before. NKT cells share features not only with Th1 and Th2 cells but also with Th17 cells. Our data provide new insights into the functional competence of NKT cells which will facilitate a better understanding of their versatile role during immune responses.
Figures
Similar articles
-
Dissociation of NKT stimulation, cytokine induction, and NK activation in vivo by the use of distinct TCR-binding ceramides.J Immunol. 2004 Jan 15;172(2):943-53. doi: 10.4049/jimmunol.172.2.943. J Immunol. 2004. PMID: 14707067
-
Regulation of immune responses by CD1d-restricted natural killer T cells.Immunol Res. 2004;30(2):139-53. doi: 10.1385/IR:30:2:139. Immunol Res. 2004. PMID: 15477656 Review.
-
IL-18 enhances IL-4 production by ligand-activated NKT lymphocytes: a pro-Th2 effect of IL-18 exerted through NKT cells.J Immunol. 2001 Jan 15;166(2):945-51. doi: 10.4049/jimmunol.166.2.945. J Immunol. 2001. PMID: 11145671
-
Molecular profiling reveals distinct functional attributes of CD1d-restricted natural killer (NK) T cell subsets.Mol Immunol. 2008 May;45(9):2607-20. doi: 10.1016/j.molimm.2007.12.022. Epub 2008 Mar 4. Mol Immunol. 2008. PMID: 18304639
-
The regulatory role of Valpha14 NKT cells in innate and acquired immune response.Annu Rev Immunol. 2003;21:483-513. doi: 10.1146/annurev.immunol.21.120601.141057. Epub 2001 Dec 19. Annu Rev Immunol. 2003. PMID: 12543936 Review.
Cited by
-
The ins and outs of type I iNKT cell development.Mol Immunol. 2019 Jan;105:116-130. doi: 10.1016/j.molimm.2018.09.023. Epub 2018 Nov 28. Mol Immunol. 2019. PMID: 30502719 Free PMC article. Review.
-
Invariant Natural Killer T-cells and their subtypes may play a role in the pathogenesis of endometriosis.Clinics (Sao Paulo). 2022 May 13;77:100032. doi: 10.1016/j.clinsp.2022.100032. eCollection 2022. Clinics (Sao Paulo). 2022. PMID: 35576870 Free PMC article.
-
Genome-wide gene expression profiling reveals unsuspected molecular alterations in pemphigus foliaceus.Immunology. 2014 Nov;143(3):381-95. doi: 10.1111/imm.12315. Immunology. 2014. PMID: 24813052 Free PMC article.
-
Effects of sodium fluoride on blood cellular and humoral immunity in mice.Oncotarget. 2017 Aug 10;8(49):85504-85515. doi: 10.18632/oncotarget.20198. eCollection 2017 Oct 17. Oncotarget. 2017. PMID: 29156736 Free PMC article.
-
Autoreactive natural killer T cells: promoting immune protection and immune tolerance through varied interactions with myeloid antigen-presenting cells.Immunology. 2010 Aug;130(4):471-83. doi: 10.1111/j.1365-2567.2010.03293.x. Epub 2010 May 11. Immunology. 2010. PMID: 20465577 Free PMC article. Review.
References
-
- Bendelac A, Lantz O, Quimby ME, Yewdell JW, Bennink JR, Brutkiewicz RR. CD1 recognition by mouse NK1+ T lymphocytes. Science. 1995;268:863–5. - PubMed
-
- Skold M, Faizunnessa NN, Wang CR, Cardell S. CD1d-specific NK1.1+ T cells with a transgenic variant TCR. J Immunol. 2000;165:168–74. - PubMed
-
- Emoto M, Kaufmann SH, Liver NKT. cells. an account of heterogeneity. Trends Immunol. 2003;24:364–9. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

