Class-specific interaction of profilin and ADF isovariants with actin in the regulation of plant development
- PMID: 17933902
- PMCID: PMC2174723
- DOI: 10.1105/tpc.107.052621
Class-specific interaction of profilin and ADF isovariants with actin in the regulation of plant development
Abstract
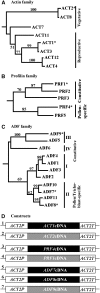
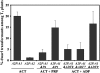
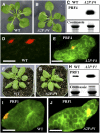
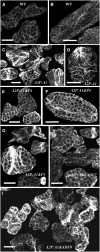
Two ancient and highly divergent actin-based cytoskeletal systems have evolved in angiosperms. Plant genomes encode complex actin and actin binding protein (ABP) gene families, most of which are phylogenetically grouped into gene classes with distinct vegetative or constitutive and reproductive expression patterns. In Arabidopsis thaliana, ectopic expression of high levels of a reproductive class actin, ACT1, in vegetative tissues causes severe dwarfing of plants with aberrant organization of most plant organs and cell types due to a severely altered actin cytoskeletal architecture. Overexpression of the vegetative class actin ACT2 to similar levels, however, produces insignificant phenotypic changes. We proposed that the misexpression of the pollen-specific ACT1 in vegetative cell types affects the dynamics of actin due to its inappropriate interaction with endogenous vegetative ABPs. To examine the functionally distinct interactions among the major classes of actins and ABPs, we ectopically coexpressed reproductive profilin (PRF4) or actin-depolymerizing factor (ADF) isovariants (e.g., ADF7) with ACT1. Our results demonstrated that the coexpression of these reproductive, but not vegetative, ABP isovariants suppressed the ectopic ACT1 expression phenotypes and restored wild-type stature and normal actin cytoskeletal architecture to the double transgenic plants. Thus, the actins and ABPs appear to have evolved class-specific, protein-protein interactions that are essential to the normal regulation of plant growth and development.
Figures




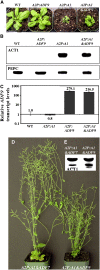
References
-
- An, Y.Q., McDowell, J.M., Huang, S., McKinney, E.C., Chambliss, S., and Meagher, R.B. (1996). Strong, constitutive expression of the Arabidopsis ACT2/ACT8 actin subclass in vegetative tissues. Plant J. 10 107–121. - PubMed
-
- Bamburg, J.R. (1999). Proteins of the ADF/cofilin family: Essential regulators of actin dynamics. Annu. Rev. Cell Dev. Biol. 15 185–230. - PubMed
-
- Carlier, M.F. (1998). Control of actin dynamics. Curr. Opin. Cell Biol. 10 45–51. - PubMed
-
- Carlsson, L., Nyström, L.-E., Sundkvist, I., Markey, F., and Lindberg, U. (1977). Actin polymerizability is influenced by profilin, a low molecular weight protein in non-muscle cells. J. Mol. Biol. 115 465–483. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

