Identification and mutational analysis of Arabidopsis FLS2 leucine-rich repeat domain residues that contribute to flagellin perception
- PMID: 17933906
- PMCID: PMC2174712
- DOI: 10.1105/tpc.106.048801
Identification and mutational analysis of Arabidopsis FLS2 leucine-rich repeat domain residues that contribute to flagellin perception
Abstract
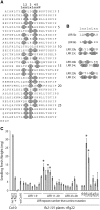
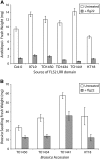
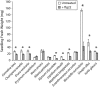
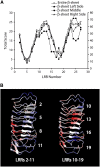
Mutational, phylogenetic, and structural modeling approaches were combined to develop a general method to study leucine-rich repeat (LRR) domains and were used to identify residues within the Arabidopsis thaliana FLAGELLIN-SENSING2 (FLS2) LRR that contribute to flagellin perception. FLS2 is a transmembrane receptor kinase that binds bacterial flagellin or a flagellin-based flg22 peptide through a presumed physical interaction within the FLS2 extracellular domain. Double-Ala scanning mutagenesis of solvent-exposed beta-strand/beta-turn residues across the FLS2 LRR domain identified LRRs 9 to 15 as contributors to flagellin responsiveness. FLS2 LRR-encoding domains from 15 Arabidopsis ecotypes and 20 diverse Brassicaceae accessions were isolated and sequenced. FLS2 is highly conserved across most Arabidopsis ecotypes, whereas more diversified functional FLS2 homologs were found in many but not all Brassicaceae accessions. flg22 responsiveness was correlated with conserved LRR regions using Conserved Functional Group software to analyze structural models of the LRR for diverse FLS2 proteins. This identified conserved spatial clusters of residues across the beta-strand/beta-turn residues of LRRs 12 to 14, the same area identified by the Ala scan, as well as other conserved sites. Site-directed randomizing mutagenesis of solvent-exposed beta-strand/beta-turn residues across LRRs 9 to 15 identified mutations that disrupt flg22 binding and showed that flagellin perception is dependent on a limited number of tightly constrained residues of LRRs 9 to 15 that make quantitative contributions to the overall phenotypic response.
Figures

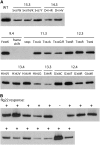
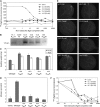
References
-
- Asai, T., Tena, G., Plotnikova, J., Willmann, M.R., Chiu, W.L., Gomez-Gomez, L., Boller, T., Ausubel, F.M., and Sheen, J. (2002). MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 415 977–983. - PubMed
-
- Ausubel, F.M., Brent, R., Kingston, R.E., Moore, D.D., Seidman, J.G., Smith, J.A., and Struhl, K. (1997). Current Protocols in Molecular Biology. (New York: John Wiley & Sons).
-
- Bauer, Z., Gomez-Gomez, L., Boller, T., and Felix, G. (2001). Sensitivity of different ecotypes and mutants of Arabidopsis thaliana toward the bacterial elicitor flagellin correlates with the presence of receptor-binding sites. J. Biol. Chem. 276 45669–45676. - PubMed
-
- Belkhadir, Y., and Chory, J. (2006). Brassinosteroid signaling: A paradigm for steroid hormone signaling from the cell surface. Science 314 1410–1411. - PubMed
-
- Bell, J.K., Mullen, G.E., Leifer, C.A., Mazzoni, A., Davies, D.R., and Segal, D.M. (2003). Leucine-rich repeats and pathogen recognition in Toll-like receptors. Trends Immunol. 24 528–533. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

