DNA indels in coding regions reveal selective constraints on protein evolution in the human lineage
- PMID: 17935613
- PMCID: PMC2151769
- DOI: 10.1186/1471-2148-7-191
DNA indels in coding regions reveal selective constraints on protein evolution in the human lineage
Abstract
Background: Insertions and deletions of DNA segments (indels) are together with substitutions the major mutational processes that generate genetic variation. Here we focus on recent DNA insertions and deletions in protein coding regions of the human genome to investigate selective constraints on indels in protein evolution.
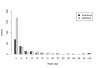
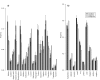
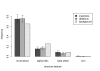
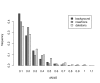
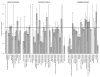
Results: Frequencies of inserted and deleted amino acids differ from background amino acid frequencies in the human proteome. Small amino acids are overrepresented, while hydrophobic, aliphatic and aromatic amino acids are strongly suppressed. Indels are found to be preferentially located in protein regions that do not form important structural domains. Amino acid insertion and deletion rates in genes associated with elementary biochemical reactions (e. g. catalytic activity, ligase activity, electron transport, or catabolic process) are lower compared to those in other genes and are therefore subject to stronger purifying selection.
Conclusion: Our analysis indicates that indels in human protein coding regions are subject to distinct levels of selective pressure with regard to their structural impact on the amino acid sequence, as well as to general properties of the genes they are located in. These findings confirm that many commonly accepted characteristics of selective constraints for substitutions are also valid for amino acid insertions and deletions.
Figures
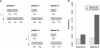
Similar articles
-
Occurrence and consequences of coding sequence insertions and deletions in Mammalian genomes.Genome Res. 2004 Apr;14(4):555-66. doi: 10.1101/gr.1977804. Genome Res. 2004. PMID: 15059996 Free PMC article.
-
Pervasive indels and their evolutionary dynamics after the fish-specific genome duplication.Mol Biol Evol. 2012 Oct;29(10):3005-22. doi: 10.1093/molbev/mss108. Epub 2012 Apr 4. Mol Biol Evol. 2012. PMID: 22490820
-
The evolution of small insertions and deletions in the coding genes of Drosophila melanogaster.Mol Biol Evol. 2013 Dec;30(12):2699-708. doi: 10.1093/molbev/mst167. Epub 2013 Sep 26. Mol Biol Evol. 2013. PMID: 24077769
-
Insertions and deletions in protein evolution and engineering.Biotechnol Adv. 2022 Nov;60:108010. doi: 10.1016/j.biotechadv.2022.108010. Epub 2022 Jun 20. Biotechnol Adv. 2022. PMID: 35738511 Review.
-
Extensive purifying selection acting on synonymous sites in HIV-1 Group M sequences.Virol J. 2008 Dec 23;5:160. doi: 10.1186/1743-422X-5-160. Virol J. 2008. PMID: 19105834 Free PMC article. Review.
Cited by
-
Correlated Selection on Amino Acid Deletion and Replacement in Mammalian Protein Sequences.J Mol Evol. 2018 Jul;86(6):365-378. doi: 10.1007/s00239-018-9853-9. Epub 2018 Jun 28. J Mol Evol. 2018. PMID: 29955898
-
Characterization of Penaeus vannamei mitogenome focusing on genetic diversity.PLoS One. 2021 Jul 30;16(7):e0255291. doi: 10.1371/journal.pone.0255291. eCollection 2021. PLoS One. 2021. PMID: 34329352 Free PMC article.
-
Computational prediction of the tolerance to amino-acid deletion in green-fluorescent protein.PLoS One. 2017 Apr 3;12(4):e0164905. doi: 10.1371/journal.pone.0164905. eCollection 2017. PLoS One. 2017. PMID: 28369116 Free PMC article.
-
Using the Evolutionary History of Proteins to Engineer Insertion-Deletion Mutants from Robust, Ancestral Templates Using Graphical Representation of Ancestral Sequence Predictions (GRASP).Methods Mol Biol. 2022;2397:85-110. doi: 10.1007/978-1-0716-1826-4_6. Methods Mol Biol. 2022. PMID: 34813061
-
Unexpectedly low mutation rates in beta-myosin heavy chain and cardiac myosin binding protein genes in Italian patients with hypertrophic cardiomyopathy.J Cell Physiol. 2011 Nov;226(11):2894-900. doi: 10.1002/jcp.22636. J Cell Physiol. 2011. PMID: 21302287 Free PMC article.
References
-
- Table of non-frameshifting indels in protein coding regions of the human genome. http://evogen.molgen.mpg.de/data/coding_indels41.txt
-
- Chen FC, Li WH. Genomic divergences between humans and other hominoids and the effective population size of the common ancestor of humans and chimpanzees. Am J Hum Genet. 2001;7(2):444–456. doi: 10.1086/318206. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pub%med&pubmedi... - DOI - PMC - PubMed
-
- Britten RJ, Rowen L, Williams J, Cameron RA. Majority of divergence between closely related DNA samples is due to indels. Proc Natl Acad Sci USA. 2003;7(8):4661–4665. doi: 10.1073/pnas.0330964100. http://dx.doi.org/10.1073/pnas.0330964100 - DOI - DOI - PMC - PubMed
-
- Watanabe H, Fujiyama A, Hattori M, Taylor TD, Toyoda A, Kuroki Y, Noguchi H, BenKahla A, Lehrach H, Sudbrak R, Kube M, Taenzer S, Galgoczy P, Platzer M, Scharfe M, Nordsiek G, Blöcker H, Hellmann I, Khaitovich P, Pääbo S, Reinhardt R, Zheng HJ, Zhang XL, Zhu GF, Wang BF, Fu G, Ren SX, Zhao GP, Chen Z, Lee YS, Cheong JE, Choi SH, Wu KM, Liu TT, Hsiao KJ, Tsai SF, Kim CG, OOta S, Kitano T, Kohara Y, Saitou N, Park HS, Wang SY, Yaspo ML, Sakaki Y. DNA sequence and comparative analysis of chimpanzee chromosome 22. Nature. 2004;7(6990):382–388. doi: 10.1038/nature02564. http://dx.doi.org/10.1038/nature02564 - DOI - DOI - PubMed
LinkOut - more resources
Full Text Sources

