Control of kidney, eye and limb expression of Bmp7 by an enhancer element highly conserved between species
- PMID: 17936743
- PMCID: PMC2394512
- DOI: 10.1016/j.ydbio.2007.08.036
Control of kidney, eye and limb expression of Bmp7 by an enhancer element highly conserved between species
Abstract
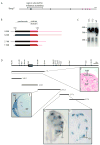
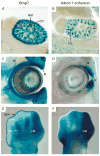
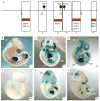
Bmp7 is expressed in numerous tissues throughout development and is required for morphogenesis of the eye, hindlimb and kidney. In this study we show that the majority if not all of the cis-regulatory sequence governing expression at these anatomical sites during development is present in approximately 20 kb surrounding exon 1. In eye, limb and kidney, multiple distinct enhancer elements drive Bmp7 expression within each organ. In the eye, the elements driving expression in the pigmented epithelium and iris are spatially separated. In the kidney, Bmp7 expression in collecting ducts and nephron progenitors is driven by separate enhancer elements. Similarly, limb mesenchyme and apical ectodermal ridge expression are governed by separate elements. Although enhancers for pigmented epithelium, nephrogenic mesenchyme and apical ectodermal ridge are distributed across the approximately 20 kb region, an element of approximately 480 base pairs within intron 1 governs expression within the developing iris, collecting duct system of the kidney and limb mesenchyme. This element is remarkably conserved both in sequence and position in the Bmp7 locus between different vertebrates, ranging from Xenopus tropicalis to Homo sapiens, demonstrating that there is strong selective pressure for Bmp7 expression at these tissue sites. Furthermore, we show that the frog enhancer functions appropriately in transgenic mice. Interestingly, the intron 1 element cannot be found in the Bmp7 genes of vertebrates such as Danio rerio and Takifugu rubripes indicating that this modification of the Bmp7 gene might have arisen during the adaptation from aquatic to terrestrial life. Mutational analysis demonstrates that the enhancer activity of the intron 1 element is entirely dependent on the presence of a 10 base pair site within the intron 1 enhancer containing a predicted binding site for the FOXD3 transcription factor.
Figures



Similar articles
-
Identification of a spatially specific enhancer element in the chicken Msx-2 gene that regulates its expression in the apical ectodermal ridge of the developing limb buds of transgenic mice.Dev Biol. 1995 Jul;170(1):230-42. doi: 10.1006/dbio.1995.1210. Dev Biol. 1995. PMID: 7601312
-
HOXA13 regulates the expression of bone morphogenetic proteins 2 and 7 to control distal limb morphogenesis.Development. 2004 Sep;131(18):4581-92. doi: 10.1242/dev.01327. Development. 2004. PMID: 15342482
-
Regulation of BMP7 expression during kidney development.Development. 1998 Sep;125(17):3473-82. doi: 10.1242/dev.125.17.3473. Development. 1998. PMID: 9693150
-
Role of BMP family members during kidney development.Int J Dev Biol. 1999;43(5):405-11. Int J Dev Biol. 1999. PMID: 10535316 Review.
-
Bone morphogenetic proteins in development.Curr Opin Genet Dev. 1996 Aug;6(4):432-8. doi: 10.1016/s0959-437x(96)80064-5. Curr Opin Genet Dev. 1996. PMID: 8791534 Review.
Cited by
-
An in vivo reporter of BMP signaling in organogenesis reveals targets in the developing kidney.BMC Dev Biol. 2008 Sep 18;8:86. doi: 10.1186/1471-213X-8-86. BMC Dev Biol. 2008. PMID: 18801194 Free PMC article.
-
Coelacanth genomes reveal signatures for evolutionary transition from water to land.Genome Res. 2013 Oct;23(10):1740-8. doi: 10.1101/gr.158105.113. Epub 2013 Jul 22. Genome Res. 2013. PMID: 23878157 Free PMC article.
-
Roles and regulation of bone morphogenetic protein-7 in kidney development and diseases.World J Stem Cells. 2016 Sep 26;8(9):288-96. doi: 10.4252/wjsc.v8.i9.288. World J Stem Cells. 2016. PMID: 27679685 Free PMC article. Review.
-
Control of the bone morphogenetic protein 7 gene in developmental and adult life.Curr Genomics. 2009 Jun;10(4):223-30. doi: 10.2174/138920209788488490. Curr Genomics. 2009. PMID: 19949543 Free PMC article.
-
A discrete transition zone organizes the topological and regulatory autonomy of the adjacent tfap2c and bmp7 genes.PLoS Genet. 2015 Jan 8;11(1):e1004897. doi: 10.1371/journal.pgen.1004897. eCollection 2015 Jan. PLoS Genet. 2015. PMID: 25569170 Free PMC article.
References
-
- Ahlberg PE, Milner AR. The origin and early diversification of tetrapods. Nature. 1994;368:507–514.
-
- Bush KT, et al. TGF-[beta] superfamily members modulate growth, branching, shaping, and patterning of the ureteric bud. Developmental Biology. 2004;266:285–298. - PubMed
-
- Dottori M, et al. The winged-helix transcription factor Foxd3 suppresses interneuron differentiation and promotes neural crest cell fate. Development. 2001;128:4127–38. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

