Genome-wide expression profiling of the Arabidopsis female gametophyte identifies families of small, secreted proteins
- PMID: 17937500
- PMCID: PMC2014789
- DOI: 10.1371/journal.pgen.0030171
Genome-wide expression profiling of the Arabidopsis female gametophyte identifies families of small, secreted proteins
Abstract
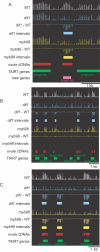
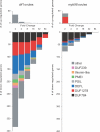
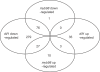
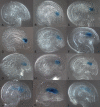
The female gametophyte of flowering plants, the embryo sac, develops within the diploid (sporophytic) tissue of the ovule. While embryo sac-expressed genes are known to be required at multiple stages of the fertilization process, the set of embryo sac-expressed genes has remained poorly defined. In particular, the set of genes responsible for mediating intracellular communication between the embryo sac and the male gametophyte, the pollen grain, is unknown. We used high-throughput cDNA sequencing and whole-genome tiling arrays to compare gene expression in wild-type ovules to that in dif1 ovules, which entirely lack embryo sacs, and myb98 ovules, which are impaired in pollen tube attraction. We identified nearly 400 genes that are downregulated in dif1 ovules. Seventy-eight percent of these embryo sac-dependent genes were predicted to encode for secreted proteins, and 60% belonged to multigenic families. Our results define a large number of candidate extracellular signaling molecules that may act during embryo sac development or fertilization; less than half of these are represented on the widely used ATH1 expression array. In particular, we found that 37 out of 40 genes encoding Domain of Unknown Function 784 (DUF784) domains require the synergid-specific transcription factor MYB98 for expression. Several DUF784 genes were transcribed in synergid cells of the embryo sac, implicating the DUF784 gene family in mediating late stages of embryo sac development or interactions with pollen tubes. The coexpression of highly similar proteins suggests a high degree of functional redundancy among embryo sac genes.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures

References
-
- Drews GN, Yadegari R. Development and function of the angiosperm female gametophyte. Annu Rev Genet. 2002;36:99–124. - PubMed
-
- Christensen CA, King JK, Jordan JR, Drews GN. Megagametogenesis in Arabidopsis wild type and the Gf mutant. Sex Plant Reprod. 1997;10:49–64.
-
- Christensen CA, Subramanian S, Drews GN. Identification of gametophytic mutations affecting female gametophyte development in Arabidopsis . Dev Biol. 1998;202:136–151. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

