Analysis on multi-domain cooperation for predicting protein-protein interactions
- PMID: 17937822
- PMCID: PMC2222654
- DOI: 10.1186/1471-2105-8-391
Analysis on multi-domain cooperation for predicting protein-protein interactions
Abstract
Background: Domains are the basic functional units of proteins. It is believed that protein-protein interactions are realized through domain interactions. Revealing multi-domain cooperation can provide deep insights into the essential mechanism of protein-protein interactions at the domain level and be further exploited to improve the accuracy of protein interaction prediction.
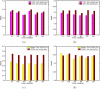
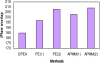
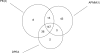
Results: In this paper, we aim to identify cooperative domains for protein interactions by extending two-domain interactions to multi-domain interactions. Based on the high-throughput experimental data from multiple organisms with different reliabilities, the interactions of domains were inferred by a Linear Programming algorithm with Multi-domain pairs (LPM) and an Association Probabilistic Method with Multi-domain pairs (APMM). Experimental results demonstrate that our approach not only can find cooperative domains effectively but also has a higher accuracy for predicting protein interaction than the existing methods. Cooperative domains, including strongly cooperative domains and superdomains, were detected from major interaction databases MIPS and DIP, and many of them were verified by physical interactions from the crystal structures of protein complexes in PDB which provide intuitive evidences for such cooperation. Comparison experiments in terms of protein/domain interaction prediction justified the benefit of considering multi-domain cooperation.
Conclusion: From the computational viewpoint, this paper gives a general framework to predict protein interactions in a more accurate manner by considering the information of both multi-domains and multiple organisms, which can also be applied to identify cooperative domains, to reconstruct large complexes and further to annotate functions of domains. Supplementary information and software are provided in http://intelligent.eic.osaka-sandai.ac.jp/chenen/MDCinfer.htm and http://zhangroup.aporc.org/bioinfo/MDCinfer.
Figures





Similar articles
-
Revealing divergent evolution, identifying circular permutations and detecting active-sites by protein structure comparison.BMC Struct Biol. 2006 Sep 2;6:18. doi: 10.1186/1472-6807-6-18. BMC Struct Biol. 2006. PMID: 16948858 Free PMC article.
-
Predicting domain-domain interaction based on domain profiles with feature selection and support vector machines.BMC Bioinformatics. 2010 Oct 29;11:537. doi: 10.1186/1471-2105-11-537. BMC Bioinformatics. 2010. PMID: 21034480 Free PMC article.
-
Yeast protein-protein interaction binding sites: prediction from the motif-motif, motif-domain and domain-domain levels.Mol Biosyst. 2010 Nov;6(11):2164-73. doi: 10.1039/c0mb00038h. Epub 2010 Aug 17. Mol Biosyst. 2010. PMID: 20714642
-
Computational prediction of protein-protein interactions.Methods Mol Biol. 2004;261:445-68. doi: 10.1385/1-59259-762-9:445. Methods Mol Biol. 2004. PMID: 15064475 Review.
-
Predicting Protein-Protein Interactions from the Molecular to the Proteome Level.Chem Rev. 2016 Apr 27;116(8):4884-909. doi: 10.1021/acs.chemrev.5b00683. Epub 2016 Apr 13. Chem Rev. 2016. PMID: 27074302 Review.
Cited by
-
DASMI: exchanging, annotating and assessing molecular interaction data.Bioinformatics. 2009 May 15;25(10):1321-8. doi: 10.1093/bioinformatics/btp142. Bioinformatics. 2009. PMID: 19420069 Free PMC article.
-
Pathway mapping and development of disease-specific biomarkers: protein-based network biomarkers.J Cell Mol Med. 2015 Feb;19(2):297-314. doi: 10.1111/jcmm.12447. Epub 2015 Jan 5. J Cell Mol Med. 2015. PMID: 25560835 Free PMC article. Review.
-
Computational prediction of the human-microbial oral interactome.BMC Syst Biol. 2014 Feb 27;8:24. doi: 10.1186/1752-0509-8-24. BMC Syst Biol. 2014. PMID: 24576332 Free PMC article.
-
DynaFace: Discrimination between Obligatory and Non-obligatory Protein-Protein Interactions Based on the Complex's Dynamics.PLoS Comput Biol. 2015 Oct 27;11(10):e1004461. doi: 10.1371/journal.pcbi.1004461. eCollection 2015 Oct. PLoS Comput Biol. 2015. PMID: 26506003 Free PMC article.
-
Proteome-wide prediction of protein-protein interactions from high-throughput data.Protein Cell. 2012 Jul;3(7):508-20. doi: 10.1007/s13238-012-2945-1. Epub 2012 Jun 22. Protein Cell. 2012. PMID: 22729399 Free PMC article. Review.
References
-
- Eisbacher M, Holmes M, Newton A, Hogg P, Khachigian L, Crossley M, Chong B. Protein-protein interaction between Fli-1 and GATA-1 mediates synergistic expression of megakaryocyte-specific genes through cooperative DNA binding. Mol Cell Biol. 2003;23:3427–3441. doi: 10.1128/MCB.23.10.3427-3441.2003. - DOI - PMC - PubMed
-
- Deng M, Sun F, Chen T. Assessment of the reliability of protein-protein interactions and protein function prediction. Pac Symp Biocomput. 2003:140–51. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

