Synergistic function of DNA methyltransferases Dnmt3a and Dnmt3b in the methylation of Oct4 and Nanog
- PMID: 17938196
- PMCID: PMC2169413
- DOI: 10.1128/MCB.01380-07
Synergistic function of DNA methyltransferases Dnmt3a and Dnmt3b in the methylation of Oct4 and Nanog
Abstract
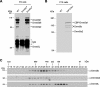
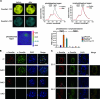
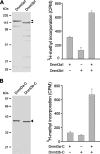
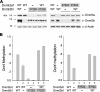
DNA methylation plays an important role in gene silencing in mammals. Two de novo methyltransferases, Dnmt3a and Dnmt3b, are required for the establishment of genomic methylation patterns in development. However, little is known about their coordinate function in the silencing of genes critical for embryonic development and how their activity is regulated. Here we show that Dnmt3a and Dnmt3b are the major components of a native complex purified from embryonic stem cells. The two enzymes directly interact and mutually stimulate each other both in vitro and in vivo. The stimulatory effect is independent of the catalytic activity of the enzyme. In differentiating embryonic carcinoma or embryonic stem cells and mouse postimplantation embryos, they function synergistically to methylate the promoters of the Oct4 and Nanog genes. Inadequate methylation caused by ablating Dnmt3a and Dnmt3b is associated with dysregulated expression of Oct4 and Nanog during the differentiation of pluripotent cells and mouse embryonic development. These results suggest that Dnmt3a and Dnmt3b form a complex through direct contact in living cells and cooperate in the methylation of the promoters of Oct4 and Nanog during cell differentiation. The physical and functional interaction between Dnmt3a and Dnmt3b represents a novel regulatory mechanism to ensure the proper establishment of genomic methylation patterns for gene silencing in development.
Figures


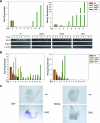
References
-
- Bachman, K. E., M. R. Rountree, and S. B. Baylin. 2001. Dnmt3a and Dnmt3b are transcriptional repressors that exhibit unique localization properties to heterochromatin. J. Biol. Chem. 276:32282-32287. - PubMed
-
- Ben-Shushan, E., E. Pikarsky, A. Klar, and Y. Bergman. 1993. Extinction of Oct-3/4 gene expression in embryonal carcinoma x fibroblast somatic cell hybrids is accompanied by changes in the methylation status, chromatin structure, and transcriptional activity of the Oct-3/4 upstream region. Mol. Cell. Biol. 13:891-901. - PMC - PubMed
-
- Bestor, T., A. Laudano, R. Mattaliano, and V. Ingram. 1988. Cloning and sequencing of a cDNA encoding DNA methyltransferase of mouse cells. The carboxyl-terminal domain of the mammalian enzymes is related to bacterial restriction methyltransferases. J. Mol. Biol. 203:971-983. - PubMed
-
- Bestor, T. H. 2000. The DNA methyltransferases of mammals. Hum. Mol. Genet. 9:2395-2402. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
