Bub1 mediates cell death in response to chromosome missegregation and acts to suppress spontaneous tumorigenesis
- PMID: 17938250
- PMCID: PMC2064762
- DOI: 10.1083/jcb.200706015
Bub1 mediates cell death in response to chromosome missegregation and acts to suppress spontaneous tumorigenesis
Abstract
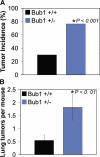
The physiological role of the mitotic checkpoint protein Bub1 is unknown. To study this role, we generated a series of mutant mice with a gradient of reduced Bub1 expression using wild-type, hypomorphic, and knockout alleles. Bub1 hypomorphic mice are viable, fertile, and overtly normal despite weakened mitotic checkpoint activity and high percentages of aneuploid cells. Bub1 haploinsufficient mice, which have a milder reduction in Bub1 protein than Bub1 hypomorphic mice, also exhibit reduced checkpoint activity and increased aneuploidy, but to a lesser extent. Although cells from Bub1 hypomorphic and haploinsufficient mice have similar rates of chromosome missegregation, cell death after an aberrant separation decreases dramatically with declining Bub1 levels. Importantly, Bub1 hypomorphic mice are highly susceptible to spontaneous tumors, whereas Bub1 haploinsufficient mice are not. These findings demonstrate that loss of Bub1 below a critical threshold drives spontaneous tumorigenesis and suggest that in addition to ensuring proper chromosome segregation, Bub1 is important for mediating cell death when chromosomes missegregate.
Figures
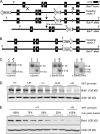
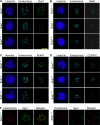

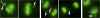
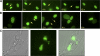

References
-
- Baker, D.J., K.B. Jeganathan, J.D. Cameron, M. Thompson, S. Juneja, A. Kopecka, R. Kumar, R.B. Jenkins, P.C. de Groen, P. Roche, and J.M. van Deursen. 2004. BubR1 insufficiency causes early onset of aging-associated phenotypes and infertility in mice. Nat. Genet. 36:744–749. - PubMed
-
- Cahill, D.P., C. Lengauer, J. Yu, G.J. Riggins, J.K. Willson, S.D. Markowitz, K.W. Kinzler, and B. Vogelstein. 1998. Mutations of mitotic checkpoint genes in human cancers. Nature. 392:300–303. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

