Identification of the cysteine nitrosylation sites in human endothelial nitric oxide synthase
- PMID: 17941803
- PMCID: PMC3893892
- DOI: 10.1089/dna.2007.0655
Identification of the cysteine nitrosylation sites in human endothelial nitric oxide synthase
Abstract
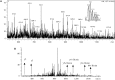
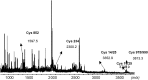
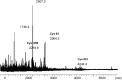
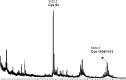
S-nitrosylation, or the replacement of the hydrogen atom in the thiol group of cysteine residues by a -NO moiety, is a physiologically important posttranslational modification. In our previous work we have shown that S-nitrosylation is involved in the disruption of the endothelial nitric oxide synthase (eNOS) dimer and that this involves the disruption of the zinc (Zn) tetrathiolate cluster due to the S-nitrosylation of Cysteine 98. However, human eNOS contains 28 other cysteine residues whose potential to undergo S-nitrosylation has not been determined. Thus, the goal of this study was to identify the cysteine residues within eNOS that are susceptible to S-nitrosylation in vitro. To accomplish this, we utilized a modified biotin switch assay. Our modification included the tryptic digestion of the S-nitrosylated eNOS protein to allow the isolation of S-nitrosylated peptides for further identification by mass spectrometry. Our data indicate that multiple cysteine residues are capable of undergoing S-nitrosylation in the presence of an excess of a nitrosylating agent. All these cysteine residues identified were found to be located on the surface of the protein according to the available X-ray structure of the oxygenase domain of eNOS. Among those identified were Cys 93 and 98, the residues involved in the formation of the eNOS dimer through a Zn tetrathiolate cluster. In addition, cysteine residues within the reductase domain were identified as undergoing S-nitrosylation. We identified cysteines 660, 801, and 1113 as capable of undergoing S-nitrosylation. These cysteines are located within regions known to bind flavin mononucleotide (FMN), flavin adenine dinucleotide (FAD), and nicotinamide adenine dinucleotide (NADPH) although from our studies their functional significance is unclear. Finally we identified cysteines 852, 975/990, and 1047/1049 as being susceptible to S-nitrosylation. These cysteines are located in regions of eNOS that have not been implicated in any known biochemical functions and the significance of their S-nitrosylation is not clear from this study. Thus, our data indicate that the eNOS protein can be S-nitrosylated at multiple sites other than within the Zn tetrathiolate cluster, suggesting that S-nitrosylation may regulate eNOS function in ways other than simply by inducing dimer collapse.
Figures


References
-
- Akashi S., Shirouza M., Terada T., Ito Y., Yokoyama S., and Takio K. (1997). Characterization of the structural difference between active and inactive forms of the ras protein by bio-chemical modification followed by mass spectrometric peptide mapping. Anal Biochem 248, 15–25 - PubMed
-
- Amado F.M.L., Dominggues P., Santana Marques M.G., Ferrer-Correia A.J., and Tomer K.B. (1997). Discrimination effects and sensitivity variations in matrix-assisted laser/ionization mass spectrometry. Rapid Commun. Mass Spectrom 11, 1347–1352
-
- Ascenzi P., Colasanti M., Persichini T., Muolo M., Polticelli F., Venturini G., Bordo D., and Bolognesi M. (2000). Re-evaluation of amino acid sequence and structural consensus rules for cysteine-nitric oxide reactivity. Biol Chem 381, 623–627 - PubMed
-
- Ascenzi P., Salvati L., Bolognesi M., Colasanti M., Polticelli F., and Venturini G. (2001). Inhibition of cysteine protease activity by NO-donors. Curr Protein Pept Sci 2, 137–153 - PubMed
-
- Belghazi M., Bathany K., Hountondji C., Grandier-Vazeille X., Manon S., and Schmitter J.-M. (2001). Analysis of protein sequences and protein complexes by matrix-assisted laser desorption/ionization mass spectrometry. Proteomics 1, 946–954 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

