The 3D rRNA modification maps database: with interactive tools for ribosome analysis
- PMID: 17947322
- PMCID: PMC2238946
- DOI: 10.1093/nar/gkm855
The 3D rRNA modification maps database: with interactive tools for ribosome analysis
Abstract
The 3D rRNA modification maps database is the first general resource of information about the locations of modified nucleotides within the 3D structure of the full ribosome, with mRNA and tRNAs in the A-, P- and E-sites. The database supports analyses for several model organisms, including higher eukaryotes, and enables users to construct 3D maps for other organisms. Data are provided for human and plant (Arabidopsis) ribosomes, and for other representative organisms from eubacteria, archaea and eukarya. Additionally, the database integrates information about positions of modifications within rRNA sequences and secondary structures, as well as links to other databases and resources about modifications and their biosynthesis. Displaying positions of modified nucleotides is fully manageable. Views of each modified nucleotide are controlled by individual buttons and buttons also control the visibility of different ribosomal molecular components. A section called 'Paint Your Own' enables the user to create a 3D modification map for rRNA from any organism where sites of modification are known. This section also provides capabilities for visualizing nucleotides of interest in rRNA or tRNA, as well as particular amino acids in ribosomal proteins. The database can be accessed at http://people.biochem.umass.edu/fournierlab/3dmodmap/
Figures

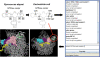
References
-
- Lapeyre B. Conserved ribosomal RNA modification and their putative roles in ribosome biogenesis and translation. In: Grosjean H, editor. Topics in Current Genetics: Fine-Tuning of RNA Functions by Modification and Editing. Vol. 12. Berlin/Heidelberg/New York: Springer-Verlag; 2005. pp. 263–284.
-
- Maden BE. The numerous modified nucleotides in eukaryotic ribosomal RNA. Prog. Nucleic Acid Res. Mol. Biol. 1990;39:241–303. - PubMed
-
- Ofengand J, Bakin A. Mapping to nucleotide resolution of pseudouridine residues in large subunit ribosomal RNAs from representative eukaryotes, prokaryotes, archaebacteria, mitochondria and chloroplasts. J. Mol. Biol. 1997;266:246–268. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

