The JmjC domain protein Epe1 prevents unregulated assembly and disassembly of heterochromatin
- PMID: 17948055
- PMCID: PMC2048757
- DOI: 10.1038/sj.emboj.7601892
The JmjC domain protein Epe1 prevents unregulated assembly and disassembly of heterochromatin
Erratum in
- EMBO J. 2008 Mar 19;27(6):921
Abstract
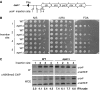
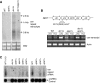
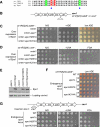
Heterochromatin normally has prescribed chromosomal positions and must not encroach on adjacent regions. We demonstrate that the fission yeast protein Epe1 stabilises silent chromatin, preventing the oscillation of heterochromatin domains. Epe1 loss leads to two contrasting phenotypes: alleviation of silencing within heterochromatin and expansion of silent chromatin into neighbouring euchromatin. Thus, we propose that Epe1 regulates heterochromatin assembly and disassembly, thereby affecting heterochromatin integrity, centromere function and chromosome segregation fidelity. Epe1 regulates the extent of heterochromatin domains at the level of chromatin, not via the RNAi pathway. Analysis of an ectopically silenced site suggests that heterochromatin oscillation occurs in the absence of heterochromatin boundaries. Epe1 requires predicted iron- and 2-oxyglutarate (2-OG)-binding residues for in vivo function, indicating that it is probably a 2-OG/Fe(II)-dependent dioxygenase. We suggest that, rather than being a histone demethylase, Epe1 may be a protein hydroxylase that affects the stability of a heterochromatin protein, or protein-protein interaction, to regulate the extent of heterochromatin domains. Thus, Epe1 ensures that heterochromatin is restricted to the domains to which it is targeted by RNAi.
Figures




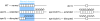
Similar articles
-
Interaction of Epe1 with the heterochromatin assembly pathway in Schizosaccharomyces pombe.Genetics. 2007 Apr;175(4):1549-60. doi: 10.1534/genetics.106.068684. Genetics. 2007. PMID: 17449867 Free PMC article.
-
Anti-silencing factor Epe1 associates with SAGA to regulate transcription within heterochromatin.Genes Dev. 2019 Jan 1;33(1-2):116-126. doi: 10.1101/gad.318030.118. Epub 2018 Dec 20. Genes Dev. 2019. PMID: 30573453 Free PMC article.
-
Regulation of ectopic heterochromatin-mediated epigenetic diversification by the JmjC family protein Epe1.PLoS Genet. 2019 Jun 17;15(6):e1008129. doi: 10.1371/journal.pgen.1008129. eCollection 2019 Jun. PLoS Genet. 2019. PMID: 31206516 Free PMC article.
-
Studies on the mechanism of RNAi-dependent heterochromatin assembly.Cold Spring Harb Symp Quant Biol. 2006;71:461-71. doi: 10.1101/sqb.2006.71.044. Cold Spring Harb Symp Quant Biol. 2006. PMID: 17381328 Review.
-
Destabilizing heterochromatin: Does Swi6/HP1 make the choice?Mol Cell. 2006 Jun 23;22(6):709-710. doi: 10.1016/j.molcel.2006.06.004. Mol Cell. 2006. PMID: 16793539 Review.
Cited by
-
Rapid epigenetic adaptation to uncontrolled heterochromatin spreading.Elife. 2015 Mar 16;4:e06179. doi: 10.7554/eLife.06179. Elife. 2015. PMID: 25774602 Free PMC article.
-
The JmjC domain of Gis1 is dispensable for transcriptional activation.FEMS Yeast Res. 2010 Nov;10(7):793-801. doi: 10.1111/j.1567-1364.2010.00680.x. Epub 2010 Sep 24. FEMS Yeast Res. 2010. PMID: 20868382 Free PMC article.
-
Set1/COMPASS repels heterochromatin invasion at euchromatic sites by disrupting Suv39/Clr4 activity and nucleosome stability.Genes Dev. 2020 Jan 1;34(1-2):99-117. doi: 10.1101/gad.328468.119. Epub 2019 Dec 5. Genes Dev. 2020. PMID: 31805521 Free PMC article.
-
Elimination of a specific histone H3K14 acetyltransferase complex bypasses the RNAi pathway to regulate pericentric heterochromatin functions.Genes Dev. 2011 Feb 1;25(3):214-9. doi: 10.1101/gad.1993611. Genes Dev. 2011. PMID: 21289066 Free PMC article.
-
Phosphorylation of Swi6/HP1 regulates transcriptional gene silencing at heterochromatin.Genes Dev. 2009 Jan 1;23(1):18-23. doi: 10.1101/gad.1708009. Genes Dev. 2009. PMID: 19136623 Free PMC article.
References
-
- Allshire RC, Javerzat JP, Redhead NJ, Cranston G (1994) Position effect variegation at fission yeast centromeres. Cell 76: 157–169 - PubMed
-
- Allshire RC, Nimmo ER, Ekwall K, Javerzat JP, Cranston G (1995) Mutations derepressing silent centromeric domains in fission yeast disrupt chromosome segregation. Genes Dev 9: 218–233 - PubMed
-
- Bannister AJ, Zegerman P, Partridge JF, Miska EA, Thomas JO, Allshire RC, Kouzarides T (2001) Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410: 120–124 - PubMed
-
- Cam HP, Sugiyama T, Chen ES, Chen X, FitzGerald PC, Grewal SI (2005) Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat Genet 37: 809–819 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

