Gateway compatible vectors for analysis of gene function in the zebrafish
- PMID: 17948311
- PMCID: PMC4518551
- DOI: 10.1002/dvdy.21354
Gateway compatible vectors for analysis of gene function in the zebrafish
Abstract
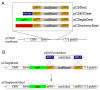
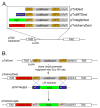
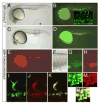
The recent establishment of recombination-based cloning systems has greatly facilitated the analysis of gene function by allowing rapid and high-efficiency generation of plasmid constructs. However, the use of such an approach in zebrafish requires the availability of recombination-compatible plasmids that are appropriate for functional studies in zebrafish embryos. In this work, we describe the construction and validation of Gateway compatible vectors based on commonly used zebrafish plasmids. We have generated pCS-based plasmids that allow rapid generation of both N-terminal and C-terminal fusion proteins, and we demonstrate that mRNA synthesized from these plasmids encodes functional native or fusion proteins in injected zebrafish embryos. In parallel, we have established similar Gateway plasmids containing Tol2 cis elements that promote efficient integration into the zebrafish genome and allow expression of native or fusion proteins in a tissue-specific manner in the zebrafish embryo. Finally, we demonstrate the use of this system to rapidly identify tissue-specific cis elements to aid the establishment of blood vessel-specific transgenic constructs. Taken together, this work provides an important platform for the rapid functional analyses of open reading frames in zebrafish embryos.
Copyright 2007 Wiley-Liss, Inc.
Figures






References
-
- Amsterdam A, Becker TS. Transgenes as screening tools to probe and manipulate the zebrafish genome. Dev Dyn. 2005;234:255–268. - PubMed
-
- Beis D, Stainier DY. In vivo cell biology: following the zebrafish trend. Trends Cell Biol. 2006;16:105–112. - PubMed
-
- Buchner DA, Su F, Yamaoka JS, Kamei M, Shavit JA, McGee B, Hanosh AW, Kim S, Jagadeeswaran P, Amigo J, Lawson ND, Goldman D, Weinstein BM, Ginsburg D, Lyons SE. From the Cover: pak2a mutations cause cerebral hemorrhage in redhead zebrafish. Proc Natl Acad Sci U S A. 2007;104:13996–14001. - PMC - PubMed
-
- Deplancke B, Mukhopadhyay A, Ao W, Elewa AM, Grove CA, Martinez NJ, Sequerra R, Doucette-Stamm L, Reece-Hoyes JS, Hope IA, Tissenbaum HA, Mango SE, Walhout AJ. A gene-centered C. elegans protein-DNA interaction network. Cell. 2006;125:1193–1205. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

